Getting Started with TissueMask
Raredon Laboratory · Yale School of Medicine
2026-04-04
Source:vignettes/getting-started.Rmd
getting-started.RmdWhy spatial masks?
In spatial transcriptomics and spatial proteomics experiments, the tissue of interest occupies an irregular region of the slide. Before computing any spatially-resolved quantity — cell-type composition, signalling gradients, diffusion fields — you need a reliable spatial mask: a polygon or multi-polygon that faithfully represents the shape of the tissue.
fit_spatial_mask() takes a set of XY point coordinates
(typically cell centroids or single-molecule transcript locations) and
fits a mask polygon to them. It returns an sf geometry
object compatible with the broader sf/GEOS/GDAL
ecosystem.
Installation
# From GitHub (requires pak or remotes)
pak::pkg_install("RaredonLab/TissueMask")
# Optional method dependencies
install.packages(c("concaveman", "MASS", "isoband", "ggplot2", "patchwork"))Method overview
method |
Topology | Speed | Key packages |
|---|---|---|---|
"raster" (default)
|
Holes + islands | Fast |
sf only |
"kde" |
Holes + islands | Moderate |
MASS, isoband
|
"concave" |
No holes | Fast | concaveman |
"convex" |
No holes | Instant |
sf only |
Shared plotting helper
We use geom_sf() throughout — not
geom_polygon(). This is essential:
geom_polygon() draws a line between consecutive rings,
producing diagonal artefacts across hole interiors.
geom_sf() delegates rendering to the sf/GEOS layer, which
handles holes correctly.
plot_mask <- function(mask, coords, title = "", subtitle = "",
fill = "#4c9be8", border = "#1a5fa8",
pt_col = "#c0392b", pt_size = 1.0, pt_alpha = 0.55) {
mask_sf <- sf::st_sf(geometry = mask)
ggplot() +
geom_sf(data = mask_sf, fill = fill, alpha = 0.2,
color = border, linewidth = 0.55) +
geom_point(data = coords, aes(x = x, y = y),
color = pt_col, size = pt_size, alpha = pt_alpha) +
coord_sf(datum = NA) +
labs(title = title, subtitle = subtitle, x = "X", y = "Y") +
theme_minimal(base_size = 10) +
theme(plot.title = element_text(face = "bold", size = 10),
plot.subtitle = element_text(size = 8, color = "grey40"),
panel.grid = element_line(color = "grey93"))
}1. Convex hull
The convex hull is the tightest convex polygon that contains all
points. It is instantaneous and requires only sf, but
cannot represent non-convex tissue shapes or internal voids.
# Simple circular cloud
coords_circle <- data.frame(x = rnorm(400, 0, 1), y = rnorm(400, 0, 1))
mask_convex <- fit_spatial_mask(coords_circle, method = "convex",
verbose = FALSE)
plot_mask(mask_convex, coords_circle,
title = "Convex hull",
subtitle = "method = 'convex'")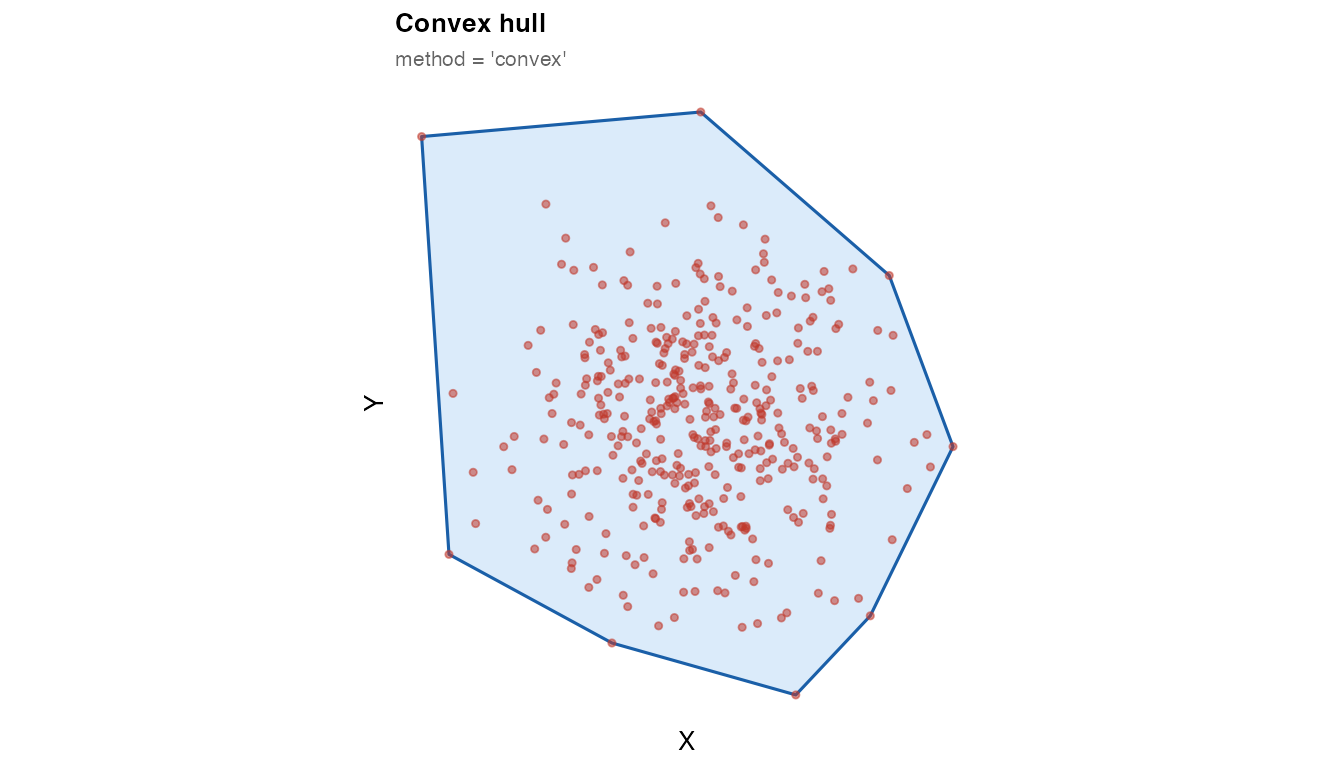
Use "convex" for quick sanity checks. For non-convex
tissues — which is the norm in practice — use "concave" or
"raster".
2. Concave hull
The concave hull wraps more tightly around the point cloud. The key
parameter is concavity: lower values produce tighter, more
concave boundaries; higher values approach the convex hull.
Requires the concaveman package.
# L-shaped cloud — non-convex; a good diagnostic
coords_L <- rbind(
data.frame(x = runif(200, 0, 5), y = runif(200, 0, 1)),
data.frame(x = runif(200, 0, 1), y = runif(200, 0, 5))
)
mask_convex_L <- fit_spatial_mask(coords_L, method = "convex",
verbose = FALSE)
mask_concave_L <- fit_spatial_mask(coords_L, method = "concave",
concavity = 1.5, verbose = FALSE)
library(patchwork)
plot_mask(mask_convex_L, coords_L,
title = "Convex hull on L-shape",
subtitle = "Incorrectly fills empty corner",
fill = "#e74c3c", border = "#922b21") +
plot_mask(mask_concave_L, coords_L,
title = "Concave hull on L-shape (concavity = 1.5)",
subtitle = "Correctly excludes the empty corner")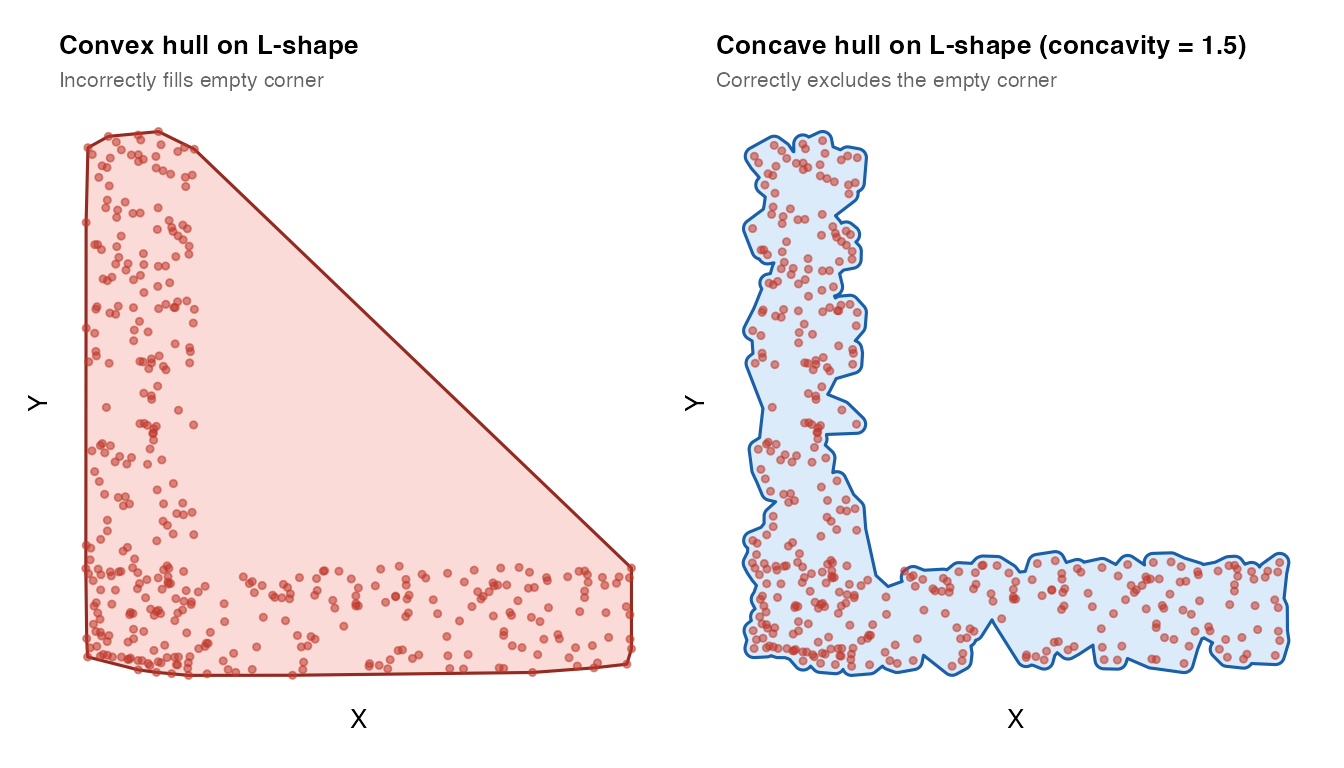
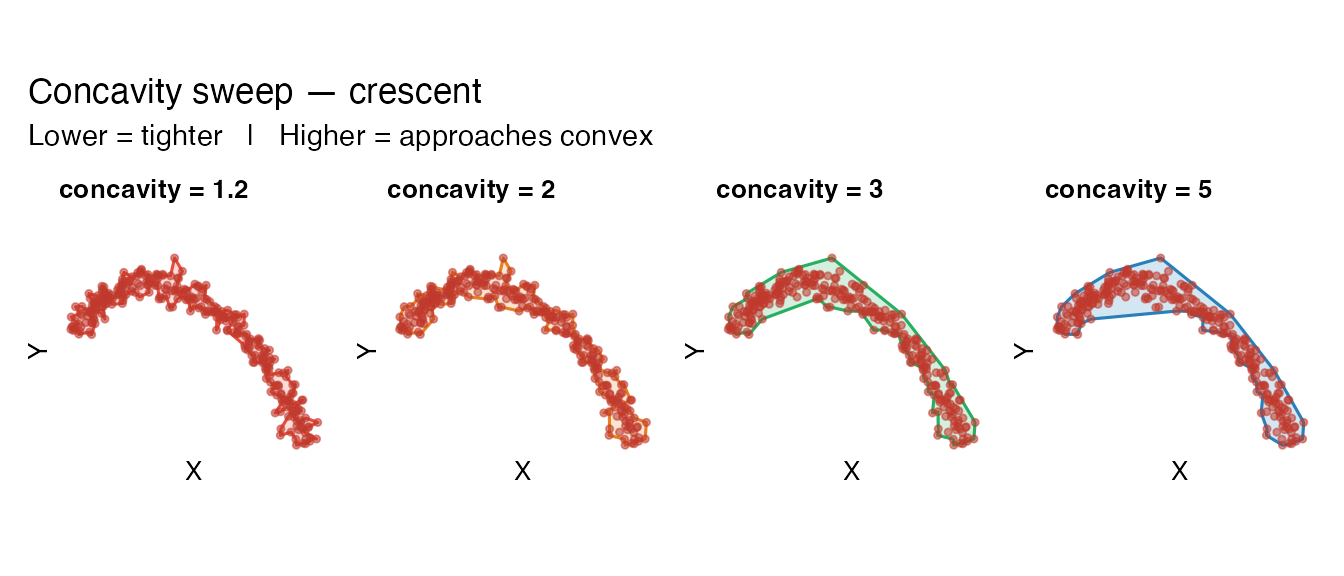
Concavity sweep
# Crescent-shaped cloud
theta <- seq(0.2, pi - 0.2, length.out = 300)
coords_cres <- data.frame(
x = cos(theta) * (1 + rnorm(300, 0, 0.08)) * 3,
y = sin(theta) * (1 + rnorm(300, 0, 0.08)) * 3 - cos(theta) * 1.5
)
pal <- c("#e74c3c", "#e67e22", "#27ae60", "#2980b9")
sw <- mapply(function(cv, col) {
m <- fit_spatial_mask(coords_cres, method = "concave",
concavity = cv, verbose = FALSE)
plot_mask(m, coords_cres, title = paste0("concavity = ", cv),
fill = col, border = col)
}, c(1.2, 2, 3, 5), pal, SIMPLIFY = FALSE)
wrap_plots(sw, nrow = 1) +
plot_annotation(
title = "Concavity sweep — crescent",
subtitle = "Lower = tighter | Higher = approaches convex"
)
3. KDE method
The KDE method estimates a 2-D kernel density surface and extracts a contour at a chosen probability level. It can represent holes and islands, but is slower for large datasets (n > 50,000). Requires MASS and isoband.
# Three-cluster cloud
mk_cluster <- function(n, cx, cy, sd = 0.4) {
data.frame(x = rnorm(n, cx, sd), y = rnorm(n, cy, sd))
}
coords_multi <- rbind(mk_cluster(150, 0, 0), mk_cluster(150, 4, 1),
mk_cluster(150, 2, 4))
mask_kde_lo <- fit_spatial_mask(coords_multi, method = "kde",
kde_threshold = 0.01, verbose = FALSE)
mask_kde_hi <- fit_spatial_mask(coords_multi, method = "kde",
kde_threshold = 0.25, verbose = FALSE)
plot_mask(mask_kde_lo, coords_multi,
title = "KDE — low threshold (0.01)",
subtitle = "One merged region",
fill = "#27ae60", border = "#1e8449") +
plot_mask(mask_kde_hi, coords_multi,
title = "KDE — high threshold (0.25)",
subtitle = "Separate masks per cluster",
fill = "#8e44ad", border = "#6c3483")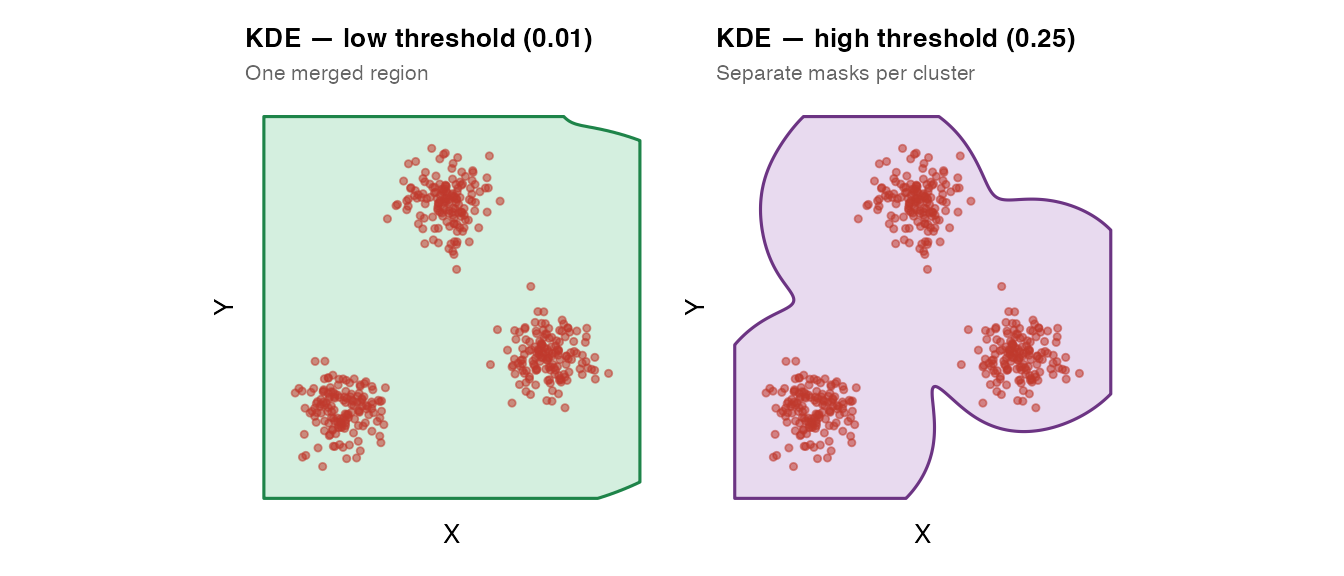
kde_threshold controls where the contour is drawn: -
Lower values → larger mask (accepts sparse regions) -
Higher values → smaller mask (only dense regions)
4. Raster method (recommended)
The raster method is the recommended default. It:
- Bins point coordinates onto a regular grid.
- Convolves the binary occupancy grid with a 2-D Gaussian of width
raster_sigma(in coordinate units — the same units asxandy). - Thresholds at
raster_threshold × max. Cells above threshold = “inside”. - Dissolves all “inside” cells via GEOS union. Holes and islands emerge automatically from the geometry — no ring-winding logic required.
- Applies a morphological close to smooth the staircase boundary.
mask_raster <- fit_spatial_mask(coords_multi, method = "raster",
verbose = FALSE)
plot_mask(mask_raster, coords_multi,
title = "Raster method (default)",
subtitle = "raster_sigma = NULL (auto = 3% of domain width)")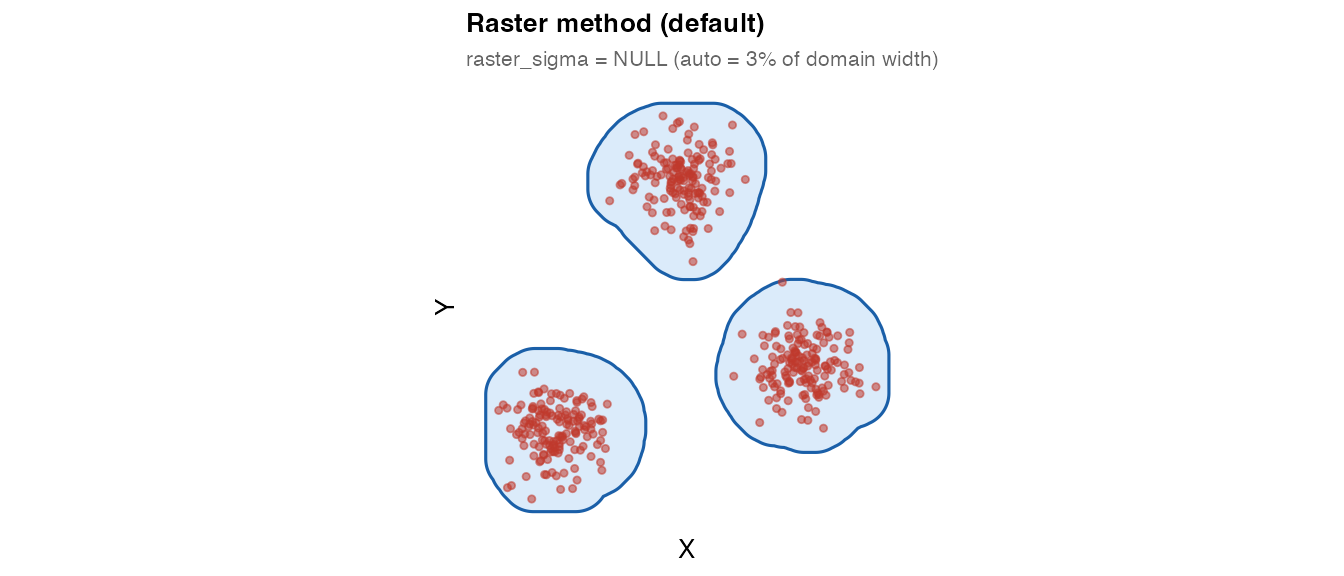
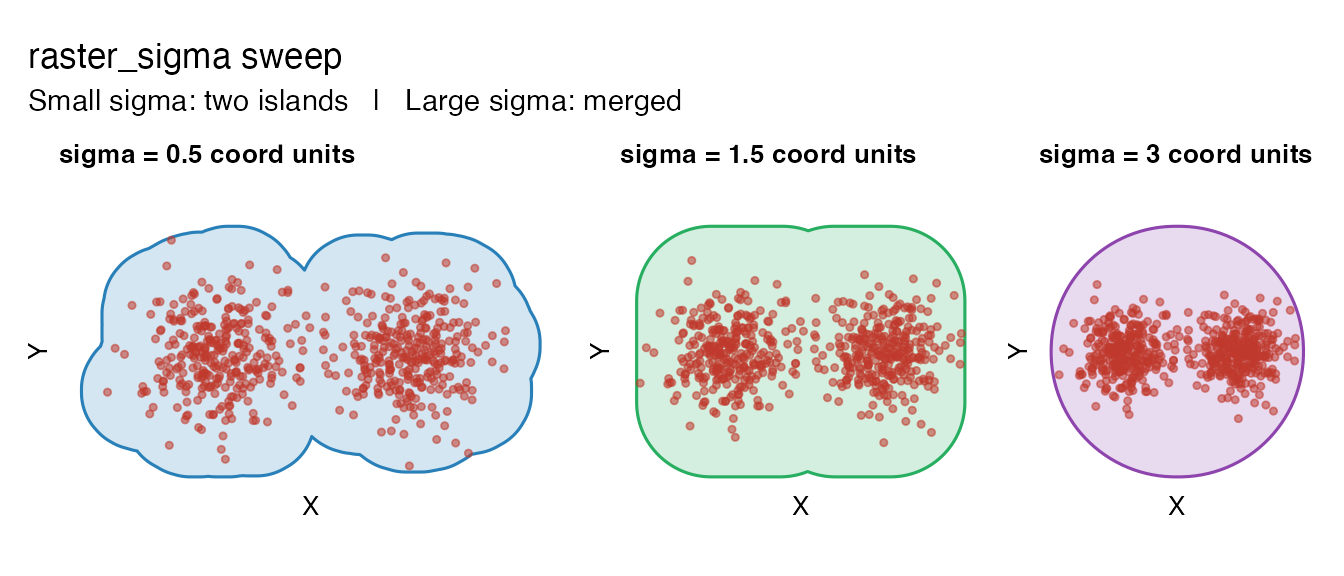
raster_sigma in coordinate units
The key innovation of the raster method is that
raster_sigma is in coordinate units, not
grid-cell units. This makes it directly interpretable: “how wide is
the Gaussian around each point?”
# Two-cluster cloud with a gap
coords_gap <- rbind(
data.frame(x = rnorm(300, 0, 1.5), y = rnorm(300, 0, 1.5)),
data.frame(x = rnorm(300, 8, 1.5), y = rnorm(300, 0, 1.5))
)
sigmas <- c(0.5, 1.5, 3.0)
pal2 <- c("#2980b9", "#27ae60", "#8e44ad")
sw2 <- mapply(function(sig, col) {
m <- fit_spatial_mask(coords_gap, method = "raster",
raster_sigma = sig, verbose = FALSE)
plot_mask(m, coords_gap,
title = paste0("sigma = ", sig, " coord units"),
fill = col, border = col)
}, sigmas, pal2, SIMPLIFY = FALSE)
wrap_plots(sw2, nrow = 1) +
plot_annotation(
title = "raster_sigma sweep",
subtitle = "Small sigma: two islands | Large sigma: merged"
)
5. Post-processing
Buffer expansion
buffer_dist expands the mask outward by a fixed distance
after fitting. Useful when you want a small margin around the tissue
edge.
mask_no_buf <- fit_spatial_mask(coords_circle, method = "raster",
buffer_dist = 0, verbose = FALSE)
mask_buf <- fit_spatial_mask(coords_circle, method = "raster",
buffer_dist = 0.4, verbose = FALSE)
plot_mask(mask_no_buf, coords_circle,
title = "No buffer (default)",
subtitle = paste0("area = ",
round(as.numeric(sf::st_area(mask_no_buf)), 2))) +
plot_mask(mask_buf, coords_circle,
title = "buffer_dist = 0.4",
subtitle = paste0("area = ",
round(as.numeric(sf::st_area(mask_buf)), 2)),
fill = "#27ae60", border = "#1e8449")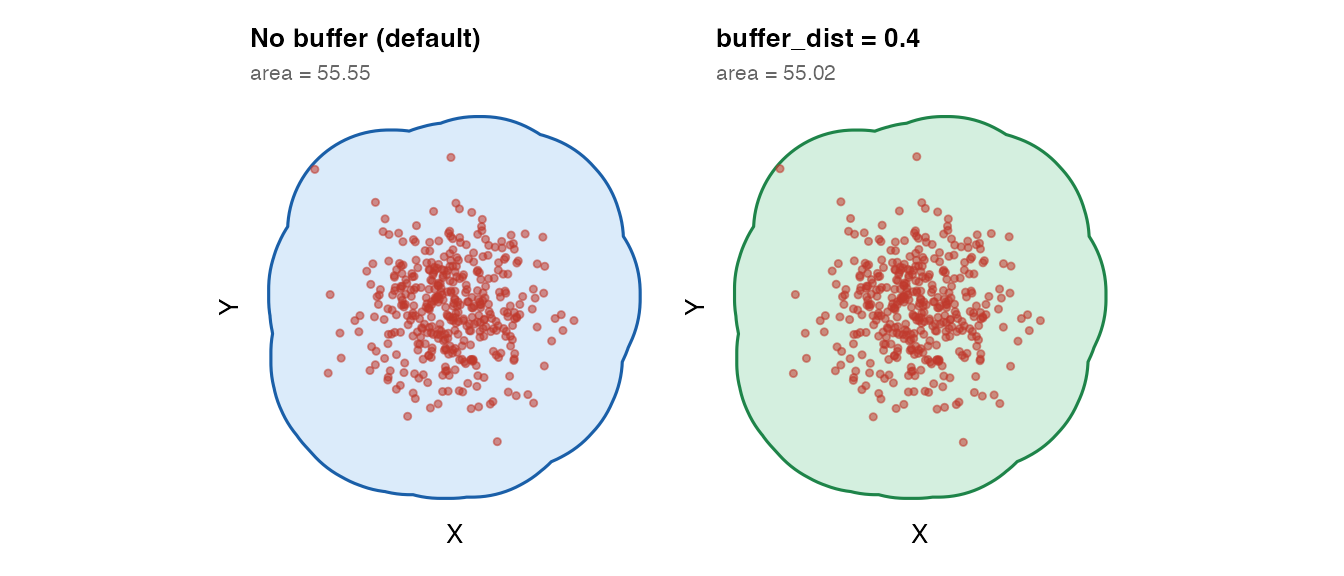
Boundary simplification
smooth_mask = TRUE applies Douglas–Peucker
simplification to reduce vertex count while preserving overall
shape.
mask_raw <- fit_spatial_mask(coords_circle, method = "raster",
smooth_mask = FALSE, verbose = FALSE)
mask_smooth <- fit_spatial_mask(coords_circle, method = "raster",
smooth_mask = TRUE,
smooth_tolerance = 0.05, verbose = FALSE)
plot_mask(mask_raw, coords_circle,
title = "No smoothing") +
plot_mask(mask_smooth, coords_circle,
title = "smooth_mask = TRUE (tolerance = 0.05)",
fill = "#8e44ad", border = "#6c3483")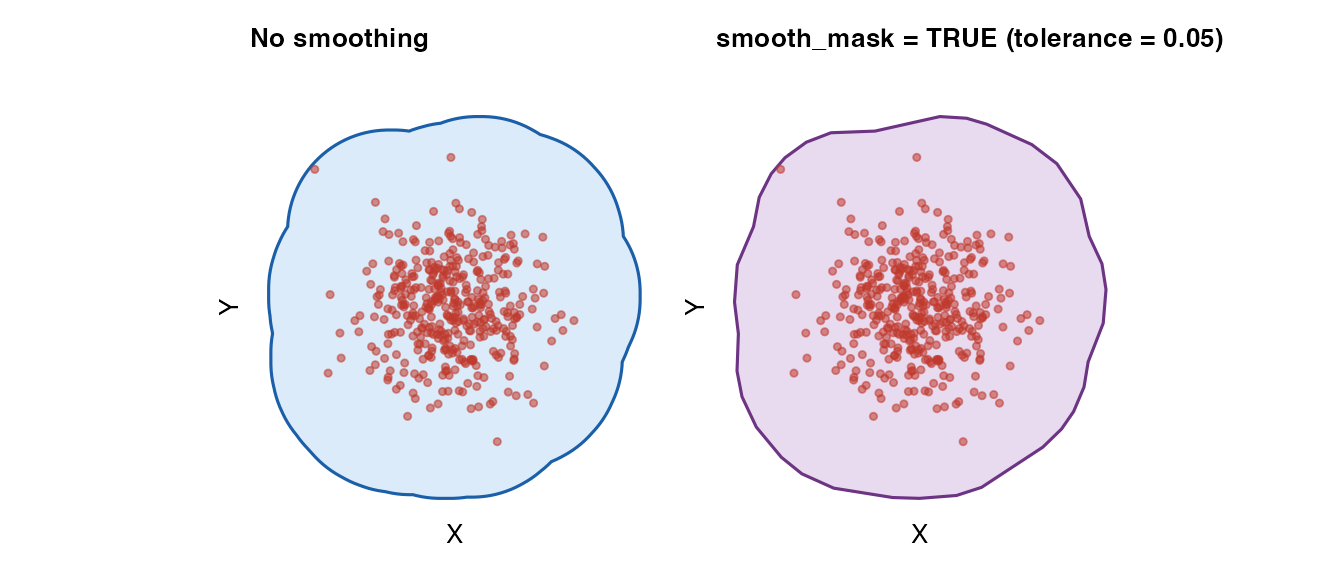
6. Verifying point containment
fit_spatial_mask() includes a built-in containment
guarantee: after fitting, all input points are checked against the mask
and a corrective buffer is applied if any fall outside. You can verify
this explicitly:
coords_test <- data.frame(
x = c(rnorm(300, 0, 3), rnorm(300, 12, 3)),
y = c(rnorm(300, 0, 3), rnorm(300, 12, 3))
)
mask_test <- fit_spatial_mask(coords_test, method = "raster", verbose = FALSE)
pts_sfc <- sf::st_as_sf(coords_test, coords = c("x", "y"),
crs = sf::NA_crs_)
# Use st_covered_by (not st_within): boundary points count as covered.
contained <- sf::st_covered_by(sf::st_geometry(pts_sfc),
sf::st_union(mask_test), sparse = FALSE)
cat("Points inside or on mask boundary:", sum(contained[, 1]),
"/", nrow(coords_test), "\n")## Points inside or on mask boundary: 600 / 6007. Method comparison
coords_demo <- rbind(
data.frame(x = rnorm(250, 0, 2), y = rnorm(250, 0, 2)),
data.frame(x = rnorm(250, 9, 2), y = rnorm(250, 9, 2))
)
m_cv <- fit_spatial_mask(coords_demo, method = "convex", verbose = FALSE)
m_cc <- fit_spatial_mask(coords_demo, method = "concave", verbose = FALSE)
m_kd <- fit_spatial_mask(coords_demo, method = "kde", verbose = FALSE)
m_ra <- fit_spatial_mask(coords_demo, method = "raster", verbose = FALSE)
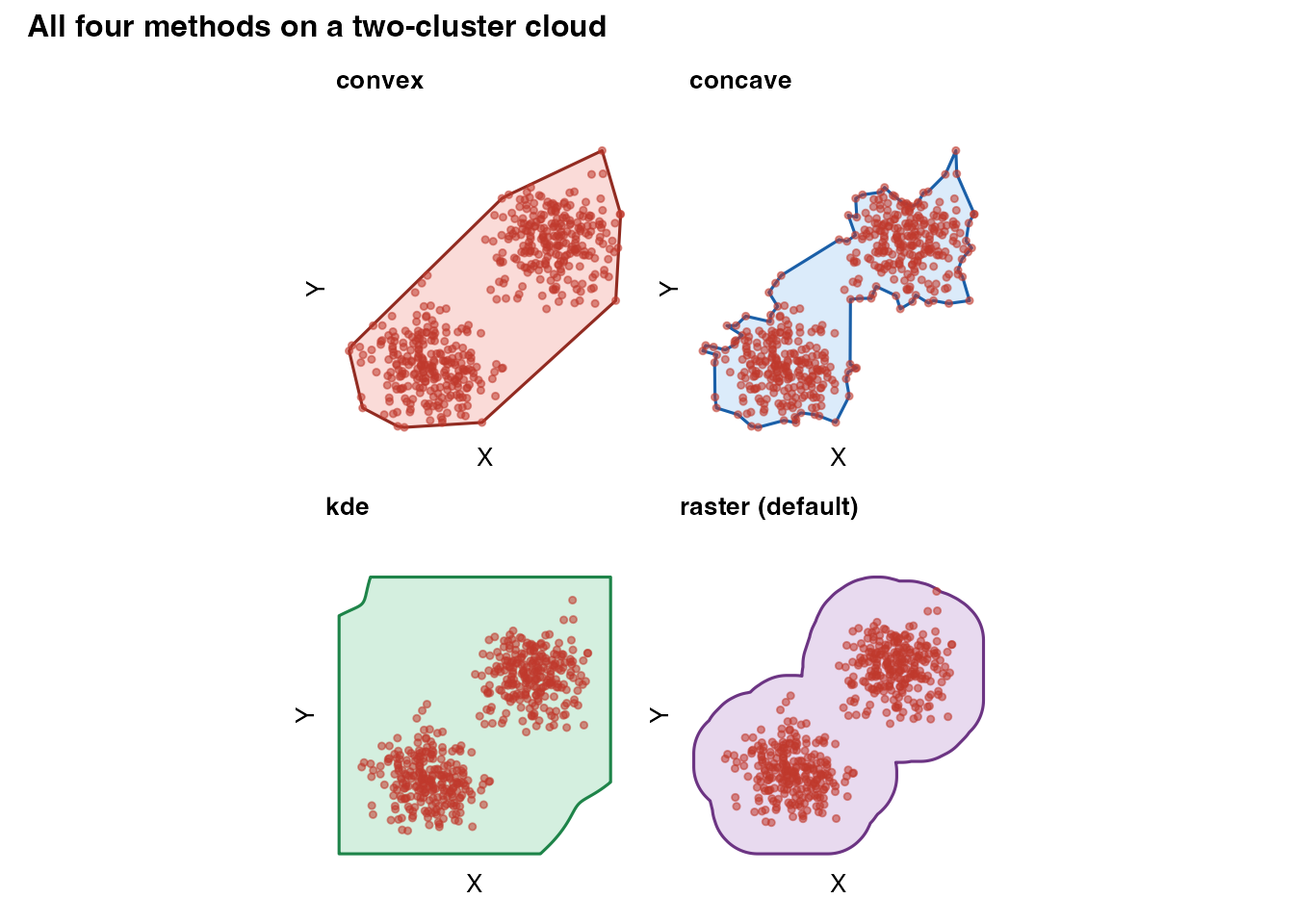
(plot_mask(m_cv, coords_demo, title = "convex",
fill = "#e74c3c", border = "#922b21") +
plot_mask(m_cc, coords_demo, title = "concave")) /
(plot_mask(m_kd, coords_demo, title = "kde",
fill = "#27ae60", border = "#1e8449") +
plot_mask(m_ra, coords_demo, title = "raster (default)",
fill = "#8e44ad", border = "#6c3483")) +
plot_annotation(
title = "All four methods on a two-cluster cloud",
theme = theme(plot.title = element_text(face = "bold", size = 12))
)
Function reference summary
fit_spatial_mask(
coords,
# Method
method = "raster", # "convex" | "concave" | "kde" | "raster"
# Concave hull
concavity = 2, # lower = tighter
length_threshold = 0,
# KDE
kde_bandwidth = NULL, # NULL = bandwidth.nrd auto
kde_threshold = 0.05, # lower = larger mask
kde_resolution = 256L,
# Raster
raster_resolution = 256L,
raster_sigma = NULL, # coordinate units; NULL = 3% of domain
raster_threshold = 0.15, # lower = larger mask
raster_min_pts = 1L,
# Post-processing
buffer_dist = 0,
smooth_mask = FALSE,
smooth_tolerance = NULL,
# Misc
n_cores = 1L,
crs = NA,
plot = FALSE,
verbose = TRUE
)Next steps
- For topologically complex tissues (holes, islands), see the Holes, Islands, and Parameter Tuning vignette.
- To estimate molecular concentration fields on a fitted mask, see the companion package TissueField.