Holes, Islands, and Parameter Tuning
Raredon Laboratory · Yale School of Medicine
2026-04-04
Source:vignettes/advanced-topology.Rmd
advanced-topology.RmdOverview
Real tissue architecture is topologically rich. Common features include:
- Vessel lumens — circular or elliptical voids inside solid tissue
- Necrotic cores — large central voids in tumour tissue
- Tissue tears and processing artefacts — irregular internal gaps
- Disconnected fragments — multiple separate tissue pieces on the same slide
The "raster" method in fit_spatial_mask()
handles all of these naturally: by dissolving above-threshold grid cells
via GEOS union, holes and islands emerge automatically from the
geometry with no special-case logic. This vignette demonstrates
each topology type and shows how to tune the two key parameters —
raster_sigma and raster_threshold — to control
the result.
1. Donut: single interior hole
The simplest topological case is a ring-shaped tissue — e.g., a cross-section through a vessel wall. Points are concentrated in an annulus; the centre is empty.
# Points uniformly distributed in an annulus (inner r=2, outer r=5)
r_d <- sqrt(runif(800, 2^2, 5^2))
th_d <- runif(800, 0, 2 * pi)
coords_donut <- data.frame(x = r_d * cos(th_d), y = r_d * sin(th_d))
# Raster: hole emerges automatically
mask_donut_raster <- fit_spatial_mask(coords_donut, method = "raster",
raster_sigma = 0.25, raster_threshold = 0.2, verbose = FALSE)
# Convex hull: fills the hole
mask_donut_convex <- fit_spatial_mask(coords_donut, method = "convex",
verbose = FALSE)
plot_mask(mask_donut_raster, coords_donut,
title = "Raster — hole detected",
subtitle = paste0("sigma = 0.25 coord units | threshold = 0.2 | ",
length(sf::st_cast(mask_donut_raster, "POLYGON")),
" polygon(s)")) +
plot_mask(mask_donut_convex, coords_donut,
title = "Convex hull — hole filled",
subtitle = "method = 'convex' cannot represent holes",
fill = "#e74c3c", border = "#922b21") +
plot_annotation(
title = "Donut: vessel lumen topology",
theme = theme(plot.title = element_text(face = "bold", size = 12))
)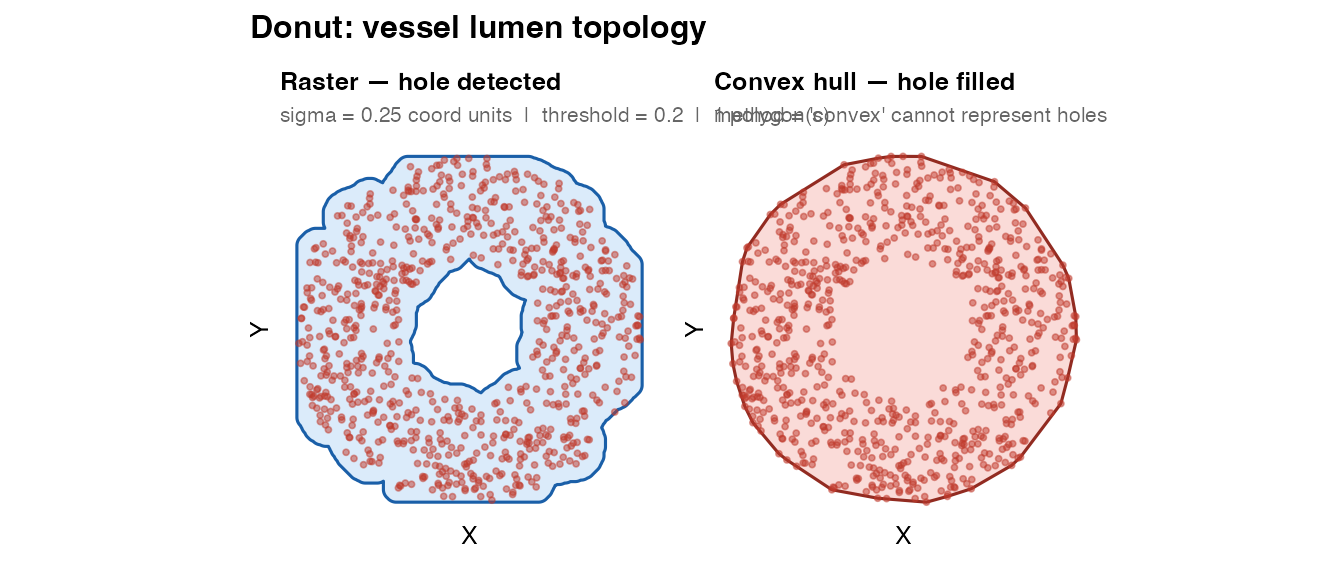
The raster mask is a POLYGON with one interior ring
(hole). The convex hull is a solid disc — it cannot represent the
void.
2. Swiss cheese: multiple holes
A square tissue block with four circular voids — representative of tissue with multiple vessel lumens or punch artefacts.
# Build ground-truth region: 10x10 square minus four circles
hdefs <- list(
list(cx = 2.5, cy = 7.5, r = 1.1),
list(cx = 7.0, cy = 6.5, r = 1.4),
list(cx = 4.0, cy = 2.5, r = 0.9),
list(cx = 8.0, cy = 2.0, r = 1.2)
)
outer_sq <- sf::st_sfc(sf::st_polygon(list(
rbind(c(0,0), c(10,0), c(10,10), c(0,10), c(0,0)))))
hcircles <- lapply(hdefs, function(h) {
sf::st_polygon(list(.circle_mat(h$cx, h$cy, h$r)))
})
swiss_region <- sf::st_difference(outer_sq,
sf::st_sfc(sf::st_union(sf::st_sfc(hcircles))))
coords_swiss <- sample_in_polygon(swiss_region, n = 1500)
mask_swiss <- fit_spatial_mask(coords_swiss, method = "raster",
raster_sigma = 0.25, raster_threshold = 0.2, verbose = FALSE)
plot_mask(mask_swiss, coords_swiss,
title = "Swiss cheese — raster method",
subtitle = "10x10 square with four circular voids")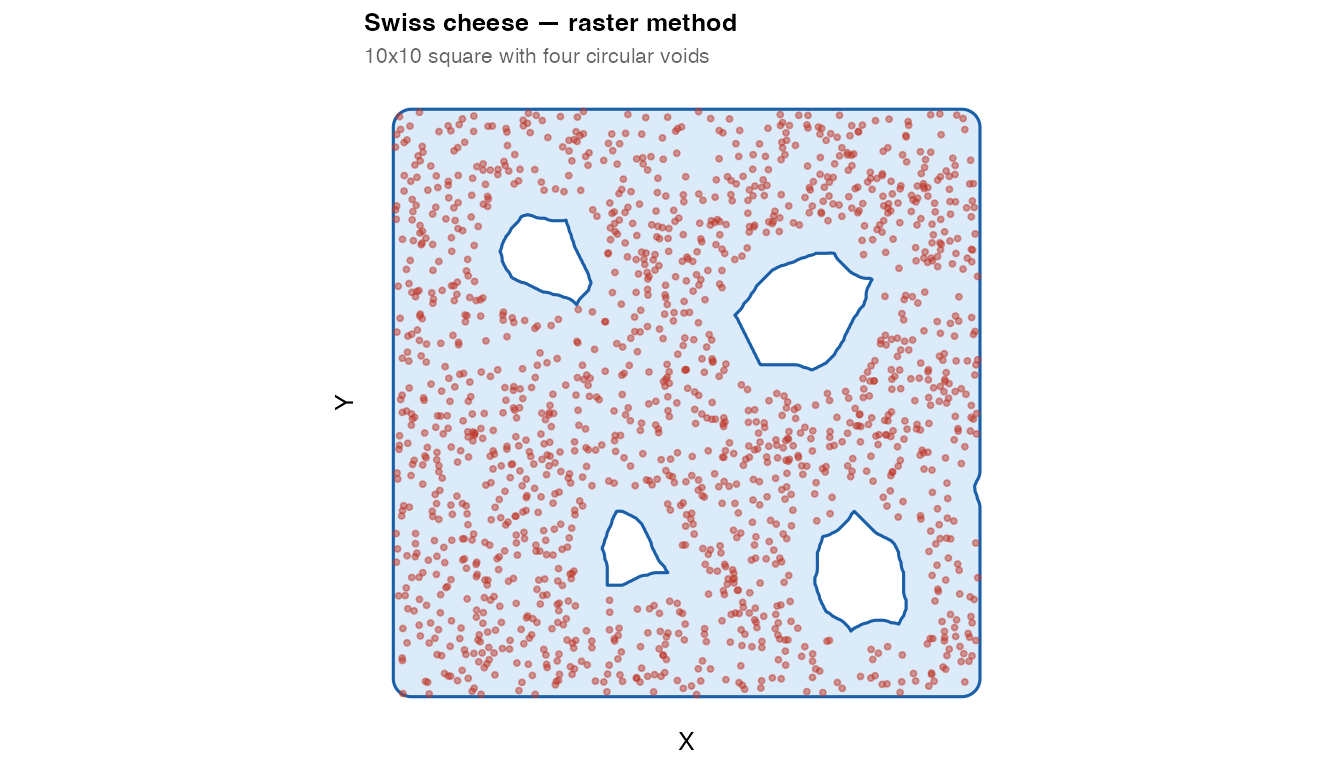
mask_swiss_concave <- fit_spatial_mask(coords_swiss, method = "concave",
concavity = 1.8, verbose = FALSE)
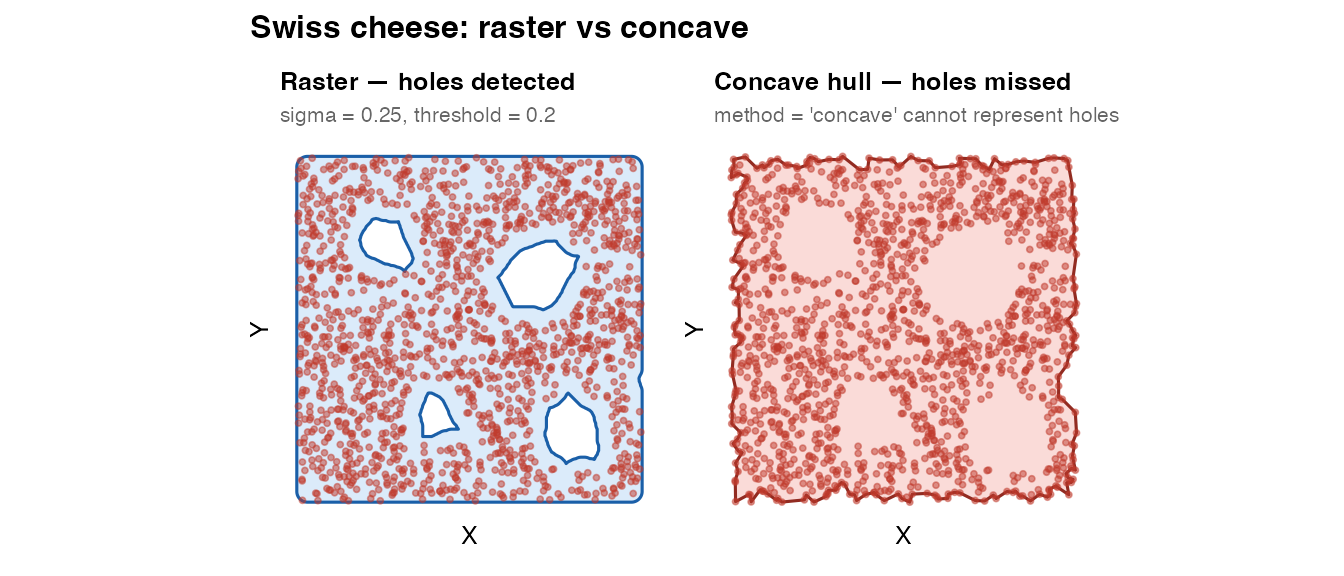
plot_mask(mask_swiss, coords_swiss,
title = "Raster — holes detected",
subtitle = "sigma = 0.25, threshold = 0.2") +
plot_mask(mask_swiss_concave, coords_swiss,
title = "Concave hull — holes missed",
subtitle = "method = 'concave' cannot represent holes",
fill = "#e74c3c", border = "#922b21") +
plot_annotation(
title = "Swiss cheese: raster vs concave",
theme = theme(plot.title = element_text(face = "bold", size = 12))
)
3. Archipelago: disconnected islands
Multiple disconnected tissue fragments — e.g., from sectioning —
appear as separate spatial clusters. The raster method represents these
as a MULTIPOLYGON, while a convex hull bridges them into
one solid region.
mk_isl <- function(n, cx, cy, rx, ry, ns = 0.15) {
th <- runif(n, 0, 2 * pi)
r <- sqrt(runif(n))
data.frame(
x = cx + r * rx * cos(th) + rnorm(n, 0, ns),
y = cy + r * ry * sin(th) + rnorm(n, 0, ns)
)
}
coords_arch <- rbind(
mk_isl(300, 0, 0, 3, 2),
mk_isl(300, 9, 4, 2, 3),
mk_isl(300, 5, -7, 2.5, 1.5)
)
mask_arch_raster <- fit_spatial_mask(coords_arch, method = "raster",
raster_sigma = 0.4, raster_threshold = 0.18, verbose = FALSE)
mask_arch_convex <- fit_spatial_mask(coords_arch, method = "convex",
verbose = FALSE)
n_islands <- length(sf::st_cast(mask_arch_raster, "POLYGON"))
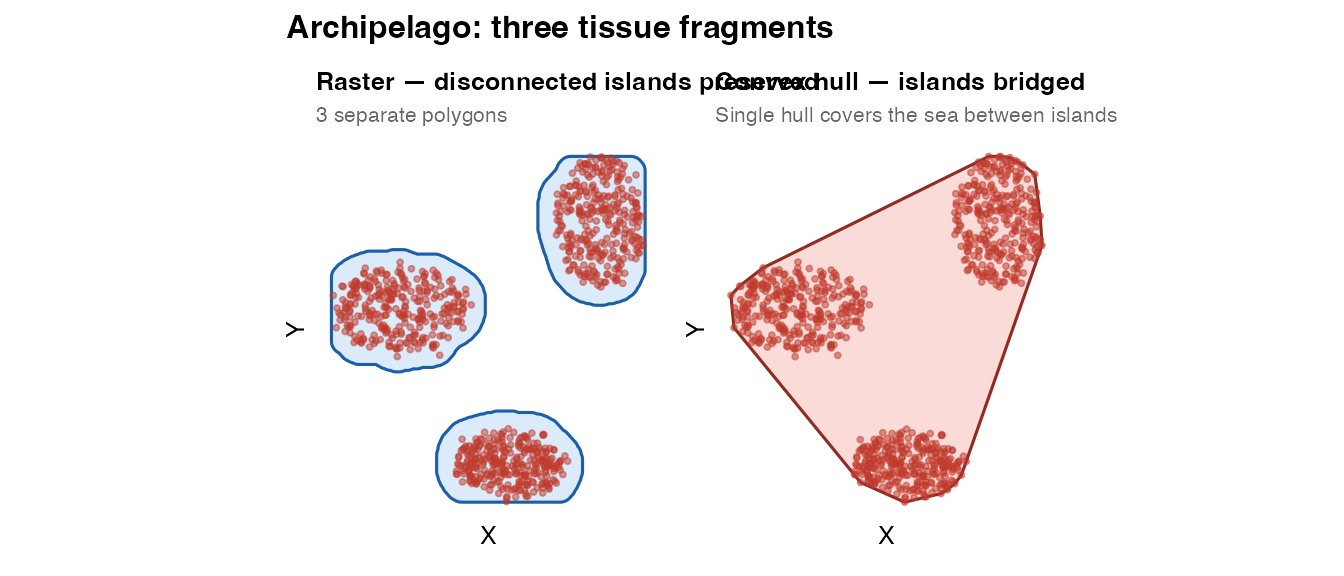
plot_mask(mask_arch_raster, coords_arch,
title = "Raster — disconnected islands preserved",
subtitle = paste0(n_islands, " separate polygons")) +
plot_mask(mask_arch_convex, coords_arch,
title = "Convex hull — islands bridged",
subtitle = "Single hull covers the sea between islands",
fill = "#e74c3c", border = "#922b21") +
plot_annotation(
title = "Archipelago: three tissue fragments",
theme = theme(plot.title = element_text(face = "bold", size = 12))
)
4. raster_sigma sweep
raster_sigma is the most important tuning parameter. It
controls the Gaussian spread applied to the occupancy grid in
coordinate units (the same units as x and
y). This makes it directly interpretable:
“How wide is the Gaussian around each point?”
-
Small
raster_sigma→ each point influences only its immediate neighbourhood → fine gaps and voids are preserved → risk of over-fragmentation -
Large
raster_sigma→ points influence a larger area → voids fill in and islands merge → risk of over-merging
# Donut cloud — clear hole with adjustable fill behaviour
sigmas <- c(0.15, 0.3, 0.6, 1.0, 1.4, 1.7)
pal <- c("#2c3e50", "#2980b9", "#27ae60",
"#f39c12", "#e74c3c", "#8e44ad")
sw_plots <- mapply(function(sig, col) {
m <- fit_spatial_mask(coords_donut, method = "raster",
raster_sigma = sig, raster_threshold = 0.2,
verbose = FALSE)
n_poly <- length(sf::st_cast(m, "POLYGON"))
plot_mask(m, coords_donut,
title = paste0("sigma = ", sig),
subtitle = paste0(n_poly, " polygon(s)"),
fill = col, border = col)
}, sigmas, pal, SIMPLIFY = FALSE)
wrap_plots(sw_plots, ncol = 3) +
plot_annotation(
title = "raster_sigma sweep on donut cloud",
subtitle = paste0("Small sigma: hole preserved | ",
"Large sigma: hole fills in"),
theme = theme(
plot.title = element_text(face = "bold", size = 12),
plot.subtitle = element_text(size = 9, color = "grey40")
)
)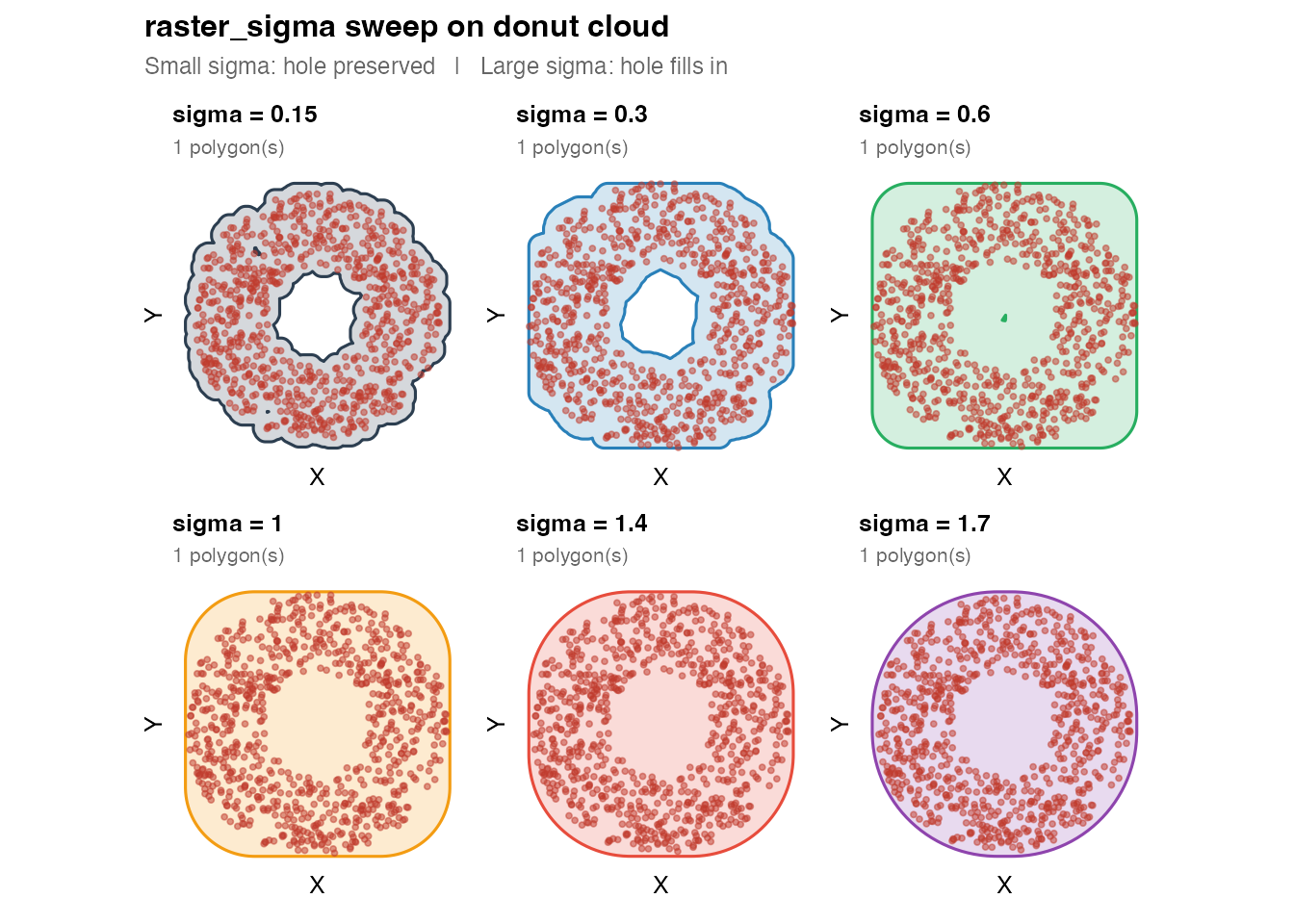
Interpretation: The hole is visible at sigma = 0.15
– 0.6. Around sigma = 1.0–1.4 it begins to fill. By sigma = 1.7 (roughly
17% of the domain width) the annulus has merged into a solid disc.
Choose raster_sigma to be just larger than the typical
inter-point gap, but smaller than the smallest void you want to
preserve.
A useful heuristic: set raster_sigma to roughly 2–3× the
typical nearest-neighbour distance in your point cloud.
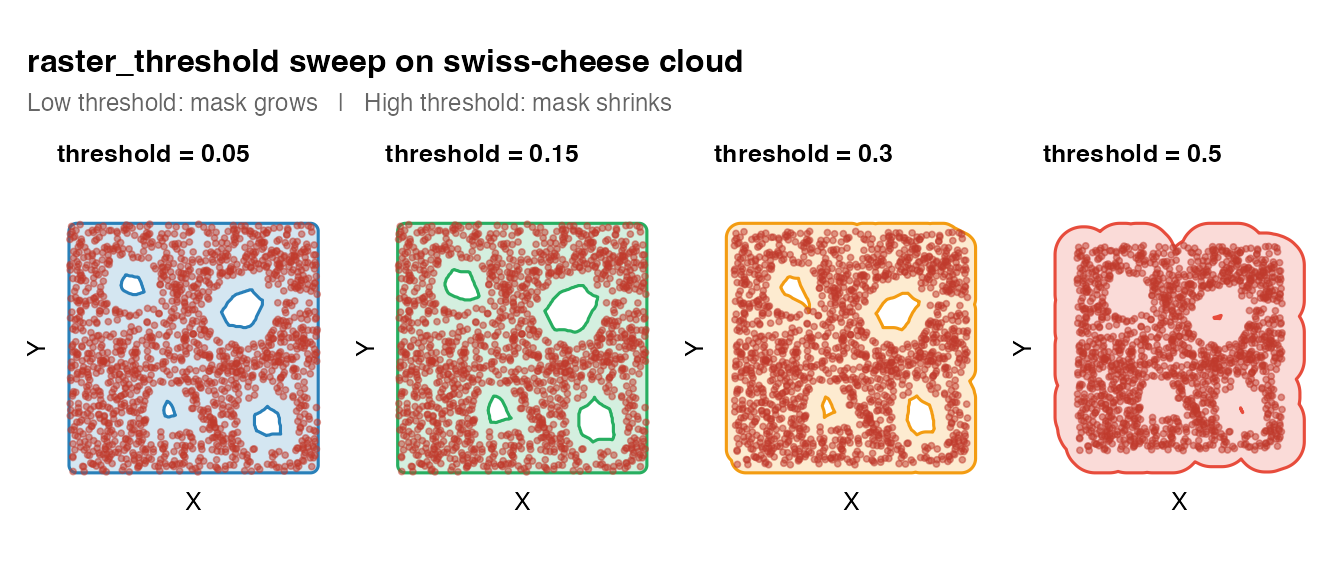
5. raster_threshold sweep
raster_threshold controls how the smoothed field is
binarised: a cell is “inside” when its smoothed value exceeds
raster_threshold × max. The effect is like adjusting the
isocontour level on a density map.
thresholds <- c(0.05, 0.15, 0.30, 0.50)
pal_th <- c("#2980b9", "#27ae60", "#f39c12", "#e74c3c")
th_plots <- mapply(function(thr, col) {
m <- fit_spatial_mask(coords_swiss, method = "raster",
raster_sigma = 0.25, raster_threshold = thr,
verbose = FALSE)
plot_mask(m, coords_swiss,
title = paste0("threshold = ", thr),
fill = col, border = col)
}, thresholds, pal_th, SIMPLIFY = FALSE)
wrap_plots(th_plots, nrow = 1) +
plot_annotation(
title = "raster_threshold sweep on swiss-cheese cloud",
subtitle = "Low threshold: mask grows | High threshold: mask shrinks",
theme = theme(
plot.title = element_text(face = "bold", size = 12),
plot.subtitle = element_text(size = 9, color = "grey40")
)
)
6. Raster vs KDE — when to choose each
# Fractured-map cloud: irregular coast + multiple interior lakes
# (small n for vignette build speed)
th_f <- seq(0, 2 * pi, length.out = 200)
r_c <- 6 + 1.5 * sin(3 * th_f) + 0.8 * sin(7 * th_f)
mat_f <- cbind(r_c * cos(th_f), r_c * sin(th_f))
mat_f <- rbind(mat_f, mat_f[1L, ])
outer_coast <- sf::st_sfc(sf::st_polygon(list(mat_f)))
make_ellipse_poly <- function(cx, cy, a, b, angle_deg, n = 60) {
th <- seq(0, 2 * pi, length.out = n)
ang <- angle_deg * pi / 180
xe <- a * cos(th); ye <- b * sin(th)
mat <- cbind(cx + xe * cos(ang) - ye * sin(ang),
cy + xe * sin(ang) + ye * cos(ang))
mat <- rbind(mat, mat[1L, ])
sf::st_polygon(list(mat))
}
lakes <- sf::st_sfc(list(
make_ellipse_poly(-2.5, 3.0, 1.5, 0.8, 30),
make_ellipse_poly( 3.5, 1.5, 1.2, 1.0, -20),
make_ellipse_poly( 1.0, -3.5, 1.8, 0.6, 60)
))
frac_region <- sf::st_difference(outer_coast, sf::st_union(lakes))
coords_frac <- sample_in_polygon(frac_region, n = 1200)
mask_frac_raster <- fit_spatial_mask(coords_frac, method = "raster",
raster_sigma = 0.5, raster_threshold = 0.18, verbose = FALSE)
mask_frac_kde <- fit_spatial_mask(coords_frac, method = "kde",
kde_resolution = 200L, kde_threshold = 0.01, verbose = FALSE)
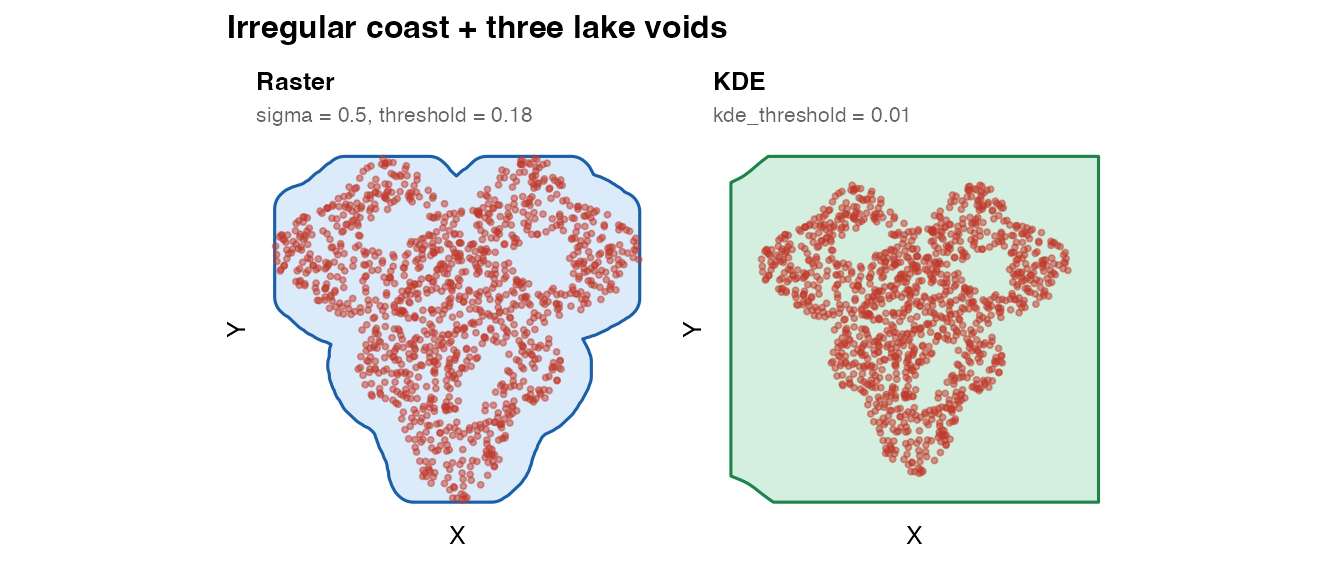
plot_mask(mask_frac_raster, coords_frac,
title = "Raster",
subtitle = "sigma = 0.5, threshold = 0.18") +
plot_mask(mask_frac_kde, coords_frac,
title = "KDE",
subtitle = "kde_threshold = 0.01",
fill = "#27ae60", border = "#1e8449") +
plot_annotation(
title = "Irregular coast + three lake voids",
theme = theme(plot.title = element_text(face = "bold", size = 12))
)
When to prefer raster: - Large point clouds (n >
50,000) — raster scales linearly; KDE is O(n²) - Datasets with sharp,
well-defined tissue boundaries - When you want explicit
raster_sigma control in interpretable units
When to prefer KDE: - Small to moderate n, dense and smooth point distributions - When bandwidth has a physical interpretation (e.g., cell radius) - When you have strongly anisotropic data and want anisotropic bandwidths
7. Scale and performance
The raster method is designed to scale to spatial transcriptomics datasets with tens or hundreds of thousands of molecules.
| n points | raster (256×256) | kde (256×256) |
|---|---|---|
| 1,000 | < 0.1 s | < 0.1 s |
| 10,000 | ~ 0.1 s | ~ 0.5 s |
| 100,000 | ~ 0.5 s | ~ 60 s |
| 1,000,000 | ~ 3 s | not recommended |
Timings approximate; hardware-dependent. KDE scales with n via
kde2d bandwidth estimation.
For very large datasets, increase raster_resolution
(default 256) to 512 for finer boundaries, at roughly 4× the compute
cost.
Parallel row-smoothing is available on Unix/macOS via
n_cores > 1:
mask <- fit_spatial_mask(
coords, method = "raster",
raster_resolution = 512L,
n_cores = 4L # Unix/macOS only
)Parameter tuning summary
| Scenario | Recommended settings |
|---|---|
| Solid tissue, no voids | Any method; "raster" default is fine |
| Vessel lumens / necrotic cores |
"raster", raster_sigma ≈ 2–3× point
spacing |
| Fine voids |
raster_sigma small (0.1–0.3× void diameter) |
| Islands should merge |
raster_sigma large (> gap width) |
| Mask too large | Increase raster_threshold (0.15 → 0.3) |
| Mask too small | Decrease raster_threshold (0.15 → 0.05) |
| Add margin around tissue | buffer_dist > 0 |
| Reduce vertex count | smooth_mask = TRUE |