Complete Reference: fit_spatial_mask()
Methods, parameters, and tuning across all use cases
Raredon Laboratory, Yale School of Medicine
2026-04-04
Source:vignettes/complete-reference.Rmd
complete-reference.RmdOverview
In spatial transcriptomics and spatial proteomics experiments, the tissue of interest occupies some irregular region of a slide. Before computing any spatially-resolved quantity (cell-type composition, signaling gradients, niche structure), you need a reliable spatial mask: a polygon or multi-polygon that faithfully represents the shape of the tissue, including internal voids (vessel lumens, necrotic cores, tissue tears) and disconnected fragments.
fit_spatial_mask() takes a set of XY point coordinates —
typically cell centroids or transcript locations — and fits a mask
polygon to them. It supports four methods of increasing complexity:
| Method | Topology | Speed | Best for |
|---|---|---|---|
"convex" |
No holes, no islands | Instant | Quick sanity checks |
"concave" |
No holes, no islands | Fast | Simple non-convex shapes |
"kde" |
Holes and islands | Moderate | Dense, smooth datasets |
"raster" |
Holes and islands | Fast at scale | Most use cases; recommended default |
Shared plotting helper
Throughout this vignette we use a single plot_mask()
helper. The key implementation note is that it uses
geom_sf() rather than geom_polygon(). This is
essential: geom_polygon() draws a line between the last
vertex of the exterior ring and the first vertex of each interior (hole)
ring, producing diagonal artefacts. geom_sf() delegates
rendering to the sf/GEOS layer which handles holes correctly.
plot_mask <- function(mask, coords, title = "", subtitle = "",
fill = "#4c9be8", border = "#1a5fa8",
pt_col = "#c0392b", pt_size = 1.2, pt_alpha = 0.55) {
mask_sf <- sf::st_sf(geometry = mask)
ggplot() +
geom_sf(data = mask_sf, fill = fill, alpha = 0.2,
color = border, linewidth = 0.55) +
geom_point(data = coords, aes(x = x, y = y),
color = pt_col, size = pt_size, alpha = pt_alpha) +
coord_sf(datum = NA) +
labs(title = title, subtitle = subtitle, x = "X", y = "Y") +
theme_minimal(base_size = 9.5) +
theme(plot.title = element_text(face = "bold", size = 9.5),
plot.subtitle = element_text(size = 8, color = "grey40"),
panel.grid = element_line(color = "grey93"))
}Function reference: fit_spatial_mask()
fit_spatial_mask(
coords,
# Method selection
method = "raster", # "convex" | "concave" | "kde" | "raster"
# Concave hull
concavity = 2, # lower = tighter; higher = approaches convex
length_threshold = 0,
# KDE
kde_bandwidth = NULL, # NULL = auto (bandwidth.nrd)
kde_threshold = 0.05, # quantile of density to threshold at
kde_resolution = 256L,
# Raster (recommended)
raster_resolution = 256L, # grid cells per axis
raster_sigma = NULL, # Gaussian spread in COORDINATE UNITS; NULL = 3% of domain
raster_threshold = 0.15, # fraction of peak Gaussian field to threshold at
raster_min_pts = 1L, # min points per cell to count as occupied
# Post-processing
buffer_dist = 0, # expand mask outward by this distance
smooth_mask = FALSE, # simplify boundary (st_simplify)
smooth_tolerance = NULL, # NULL = 1% of bounding box diagonal
# Misc
n_cores = 1L,
crs = NA,
plot = FALSE,
verbose = TRUE
)Return value
fit_spatial_mask() returns an sfc geometry
object (the sf package’s geometry column class). You
can:
- Pass it directly to
estimate_concentration_field()as themaskargument - Plot it with
geom_sf() - Compute area with
sf::st_area() - Test point containment with
sf::st_covered_by() - Convert to a data frame with
as.data.frame(sf::st_coordinates(mask))
Section 1 — Circular cloud: sanity check
The simplest possible test: a Gaussian cloud with no special topology. All three non-trivial methods should return similar results here. This section establishes that the function runs and that the three main methods agree on simple inputs.
n <- 400
coords_circle <- data.frame(x = rnorm(n, 0, 1),
y = rnorm(n, 0, 1))
m1a <- fit_spatial_mask(coords_circle, method = "convex", verbose = FALSE)
m1b <- fit_spatial_mask(coords_circle, method = "concave", concavity = 2,
verbose = FALSE)
m1c <- fit_spatial_mask(coords_circle, method = "kde",
kde_threshold = 0.02, buffer_dist = 0.1,
verbose = FALSE)
plot_mask(m1a, coords_circle, title = "1a. Convex hull",
subtitle = "method='convex'") +
plot_mask(m1b, coords_circle, title = "1b. Concave hull",
subtitle = "concavity=2") +
plot_mask(m1c, coords_circle, title = "1c. KDE contour",
subtitle = "kde_threshold=0.02, buffer=0.1",
fill = "#27ae60", border = "#1e8449") +
plot_annotation(
title = "Section 1: Circular Cloud -- Sanity Check",
theme = theme(plot.title = element_text(face = "bold", size = 13)))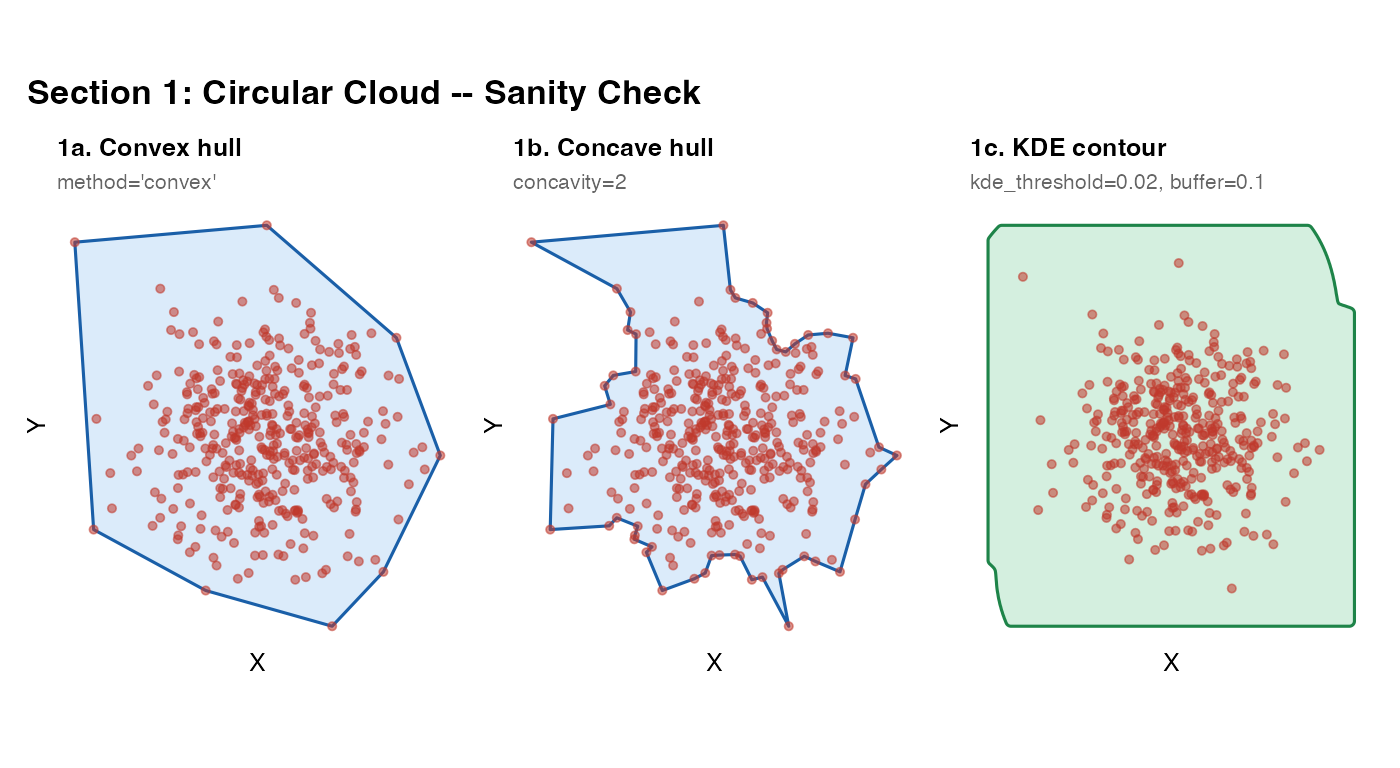
What to look for: All three masks should closely envelope the point cloud. Small differences near the boundary are expected – convex gives a polygon, KDE gives a smoother contour. Any large discrepancy here indicates a parameter problem.
Section 2 – Elongated cloud: anisotropy
Elongated tissue (e.g. a muscle section, a linear airway epithelium) has very different extents along its two axes. All methods should handle this naturally; this section confirms that no method is implicitly assuming isotropy.
coords_elong <- data.frame(x = rnorm(n, 0, 3),
y = rnorm(n, 0, 0.6))
m2a <- fit_spatial_mask(coords_elong, method = "convex", verbose = FALSE)
m2b <- fit_spatial_mask(coords_elong, method = "concave", verbose = FALSE)
m2c <- fit_spatial_mask(coords_elong, method = "kde",
kde_threshold = 0.02, buffer_dist = 0.1,
verbose = FALSE)
plot_mask(m2a, coords_elong, title = "2a. Convex") +
plot_mask(m2b, coords_elong, title = "2b. Concave") +
plot_mask(m2c, coords_elong, title = "2c. KDE",
fill = "#27ae60", border = "#1e8449") +
plot_annotation(
title = "Section 2: Elongated Cloud",
theme = theme(plot.title = element_text(face = "bold", size = 13)))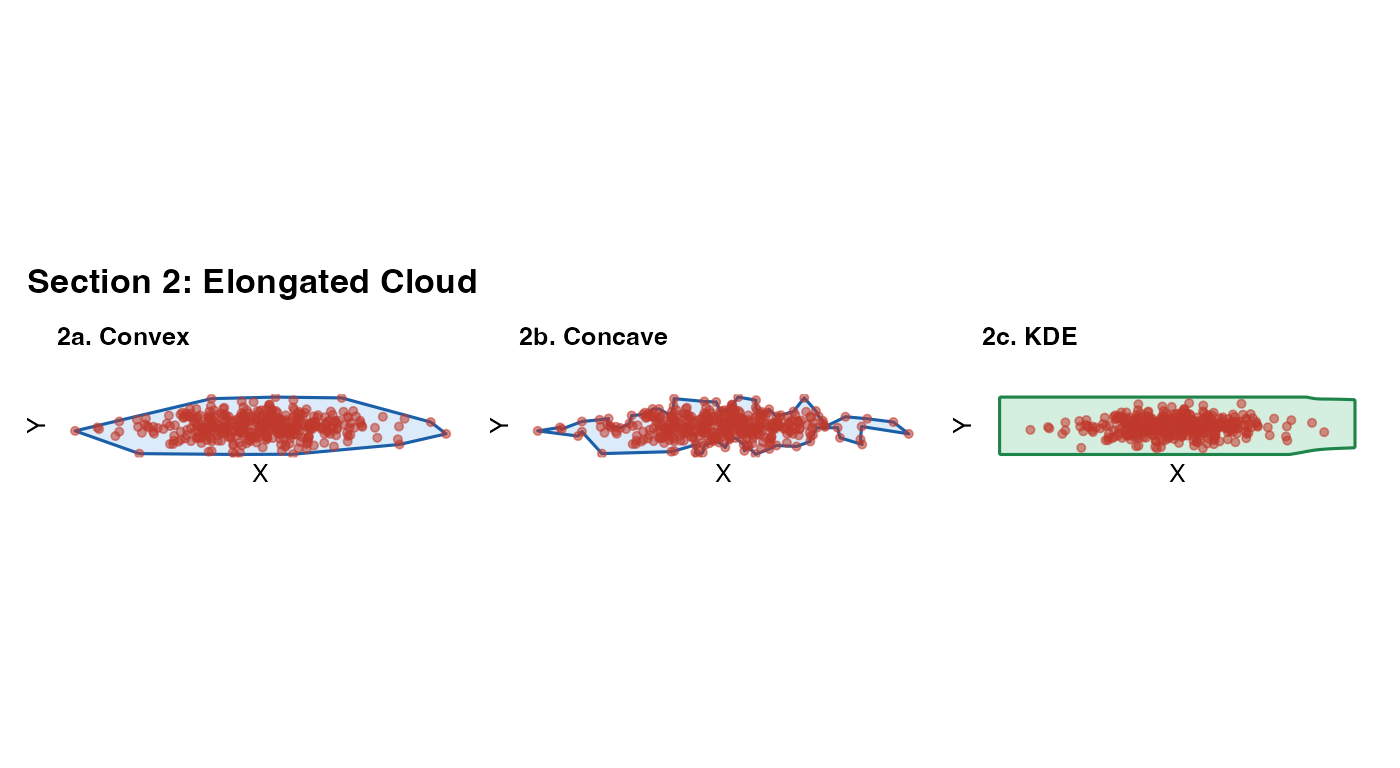
What to look for: The mask should track the long
axis of the cloud, not default to a circle. The KDE bandwidth is
computed separately for x and y (MASS::bandwidth.nrd), so
it adapts to anisotropy automatically.
Section 3 – L-shaped cloud: the non-convex diagnostic
This is the most important basic test. The L-shape is the canonical non-convex case: the convex hull incorrectly fills the concave corner of the L, including empty space that contains no tissue. A good mask must track the actual boundary and exclude the corner.
Rule of thumb: If your tissue has any re-entrant angles (C-shapes, crescents, airways, arcs), do not use
method = "convex". Usemethod = "concave"ormethod = "raster".
coords_L <- rbind(
data.frame(x = runif(200, 0, 5) + rnorm(200, 0, 0.05),
y = runif(200, 0, 1) + rnorm(200, 0, 0.05)), # horizontal bar
data.frame(x = runif(200, 0, 1) + rnorm(200, 0, 0.05),
y = runif(200, 0, 5) + rnorm(200, 0, 0.05))) # vertical bar
m3a <- fit_spatial_mask(coords_L, method = "convex", verbose = FALSE)
m3b <- fit_spatial_mask(coords_L, method = "concave", concavity = 1.5, verbose = FALSE)
m3c <- fit_spatial_mask(coords_L, method = "concave", concavity = 3.0, verbose = FALSE)
plot_mask(m3a, coords_L,
title = "3a. Convex hull",
subtitle = "Incorrectly includes empty corner",
fill = "#e74c3c", border = "#922b21") +
plot_mask(m3b, coords_L,
title = "3b. Concave, tight (1.5)",
subtitle = "Correctly excludes corner") +
plot_mask(m3c, coords_L,
title = "3c. Concave, relaxed (3.0)",
subtitle = "Partially includes corner",
fill = "#8e44ad", border = "#6c3483") +
plot_annotation(
title = "Section 3: L-Shaped Cloud -- Non-Convex",
subtitle = "Lower concavity wraps more tightly; convex hull fails",
theme = theme(plot.title = element_text(face = "bold", size = 13),
plot.subtitle = element_text(size = 9, color = "grey40")))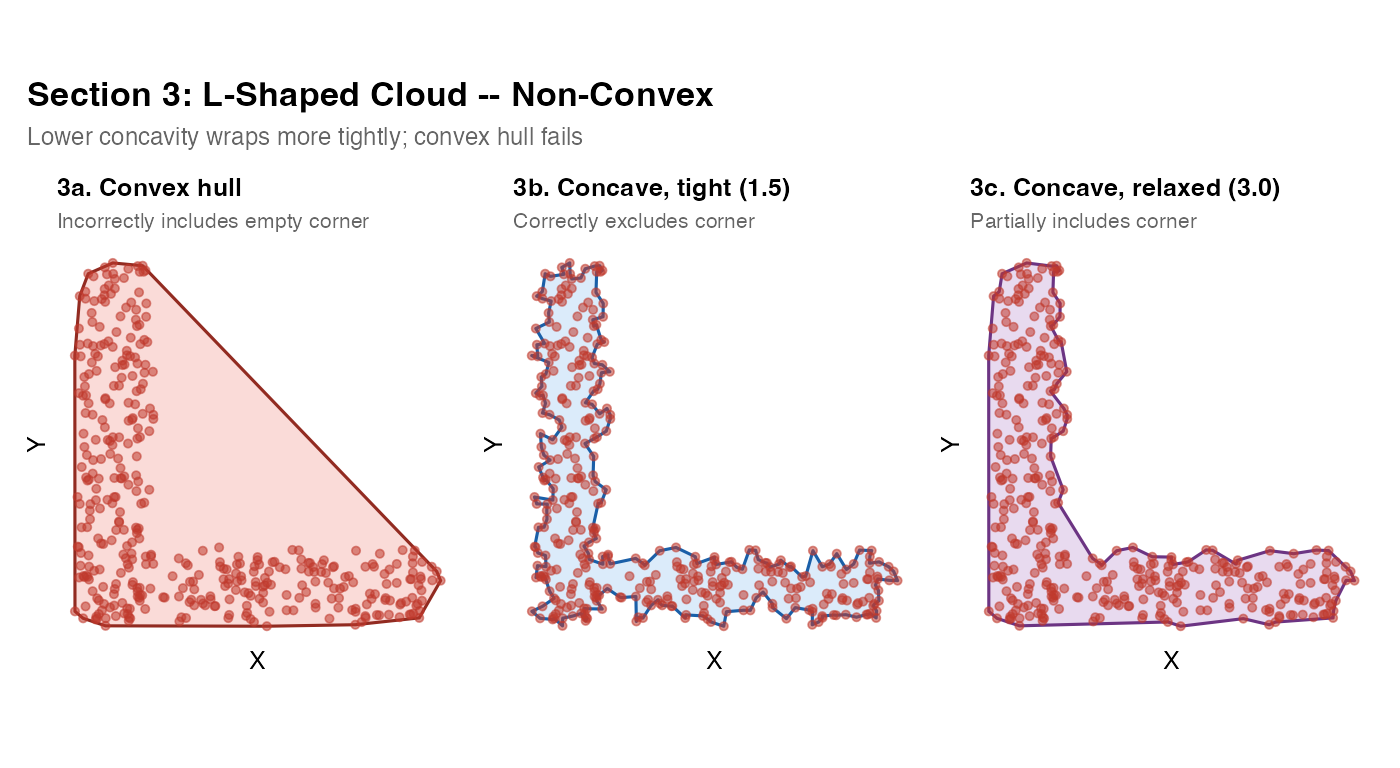
The concavity parameter
concavity controls how aggressively the hull “cuts in”
to follow the data boundary. It is the key tuning parameter for
method = "concave".
| Value | Behaviour |
|---|---|
| 1.0–1.5 | Very tight – tracks the data boundary closely |
| 2.0 | Default – good balance for most tissue shapes |
| 3.0–5.0 | Relaxed – approaches the convex hull |
When to use lower values: C-shapes, crescents,
U-shapes, any tissue with a notch or bay. When to use higher
values: Dense blobs where you want a smooth, padded boundary.
If you see the mask cutting through the interior of your tissue,
increase concavity. If it fills in empty space at
concavities, decrease it.
Section 4 – Multi-cluster blobs: connected vs. isolated masks
When tissue fragments are well-separated, a low
kde_threshold will produce a single mask that bridges them
(because the KDE density never drops to zero between clusters at low
thresholds). A higher threshold separates them into distinct
polygons.
The raster method’s raster_sigma plays the analogous
role: a large sigma bridges clusters; a small sigma isolates them.
mk_cluster <- function(n, cx, cy, sd = 0.4)
data.frame(x = rnorm(n, cx, sd), y = rnorm(n, cy, sd))
coords_multi <- rbind(mk_cluster(150, 0, 0),
mk_cluster(150, 4, 1),
mk_cluster(150, 2, 4))
m4a <- fit_spatial_mask(coords_multi, method = "convex", verbose = FALSE)
m4b <- fit_spatial_mask(coords_multi, method = "concave", concavity = 2,
verbose = FALSE)
m4c <- fit_spatial_mask(coords_multi, method = "kde",
kde_threshold = 0.01, buffer_dist = 0.15,
verbose = FALSE)
m4d <- fit_spatial_mask(coords_multi, method = "kde",
kde_threshold = 0.25, buffer_dist = 0.15,
verbose = FALSE)
(plot_mask(m4a, coords_multi, title = "4a. Convex") +
plot_mask(m4b, coords_multi, title = "4b. Concave")) /
(plot_mask(m4c, coords_multi,
title = "4c. KDE -- inclusive",
subtitle = "threshold=0.01: merged",
fill = "#27ae60", border = "#1e8449") +
plot_mask(m4d, coords_multi,
title = "4d. KDE -- tight",
subtitle = "threshold=0.25: separated",
fill = "#8e44ad", border = "#6c3483")) +
plot_annotation(
title = "Section 4: Multi-Cluster",
theme = theme(plot.title = element_text(face = "bold", size = 13)))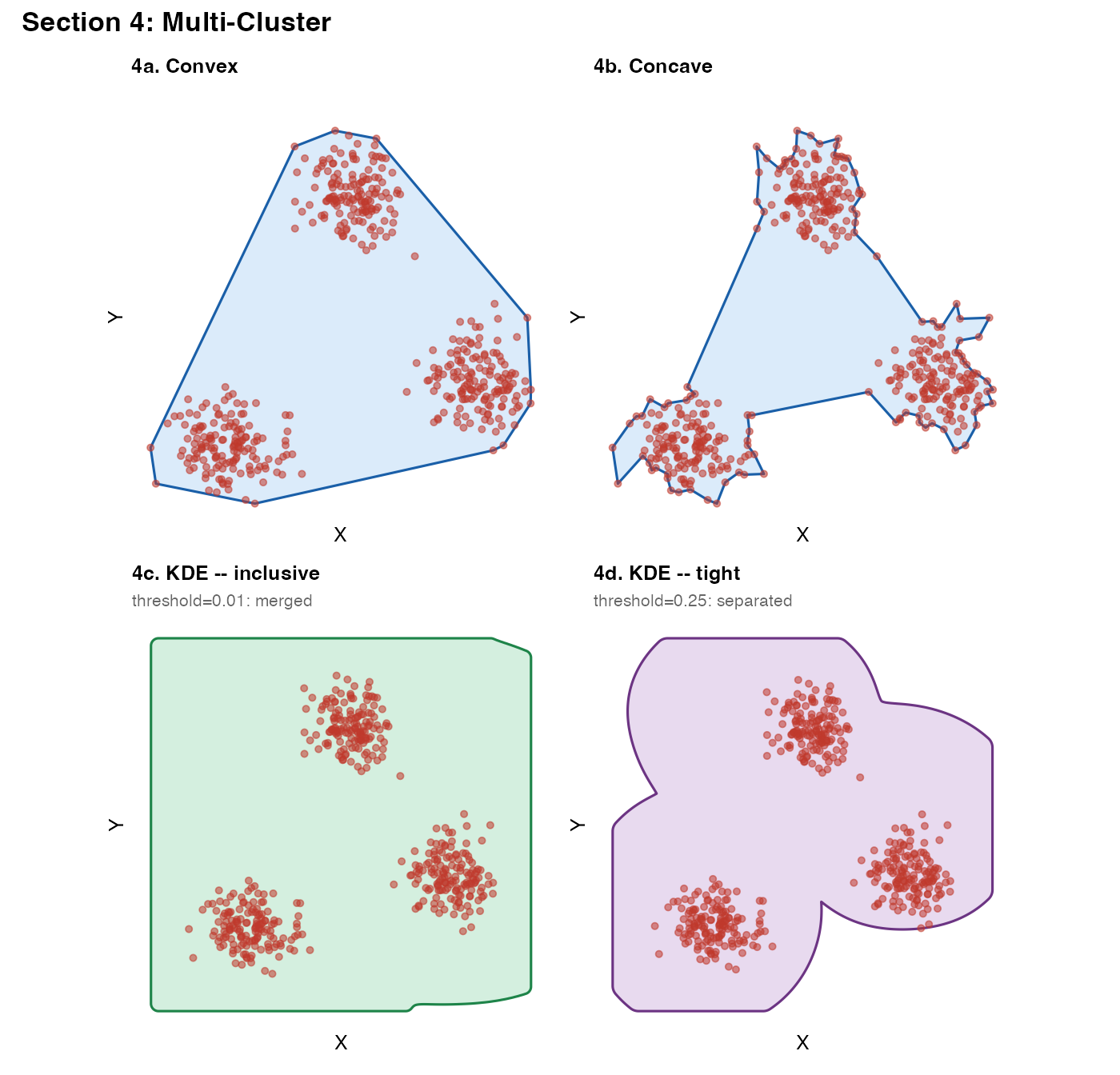
Choosing kde_threshold
kde_threshold is expressed as a quantile of the KDE
density values across the grid.
- Low values (0.01–0.05): the contour is drawn far out in the low-density tails. Clusters merge; mask is generous.
- High values (0.15–0.30): only the dense core is enclosed. Clusters separate; mask is tight.
Practical advice: Start at 0.05 and increase until the mask matches your visual inspection. If you expect biologically separated regions (e.g. two lobes), a threshold that separates them is usually correct.
Section 5 – Concavity sweep on a crescent
A crescent is a severe test of the concave hull: it has a large, deep
concavity spanning most of the shape. This sweep lets you calibrate the
concavity parameter on a shape where the expected correct
answer is obvious.
theta <- seq(0.2, pi - 0.2, length.out = 300)
coords_cres <- data.frame(
x = cos(theta) * (1 + rnorm(300, 0, 0.08)) * 3,
y = sin(theta) * (1 + rnorm(300, 0, 0.08)) * 3 - cos(theta) * 1.5)
pal <- c("#e74c3c", "#e67e22", "#27ae60", "#2980b9")
sw <- mapply(function(cv, col) {
m <- fit_spatial_mask(coords_cres, method = "concave",
concavity = cv, verbose = FALSE)
plot_mask(m, coords_cres, title = paste0("concavity = ", cv),
fill = col, border = col)
}, c(1.2, 2, 3, 5), pal, SIMPLIFY = FALSE)
wrap_plots(sw, nrow = 1) +
plot_annotation(
title = "Section 5: Concavity Sweep -- Crescent",
subtitle = "Lower = tighter | Higher = approaches convex hull",
theme = theme(plot.title = element_text(face = "bold", size = 13),
plot.subtitle = element_text(size = 9, color = "grey40")))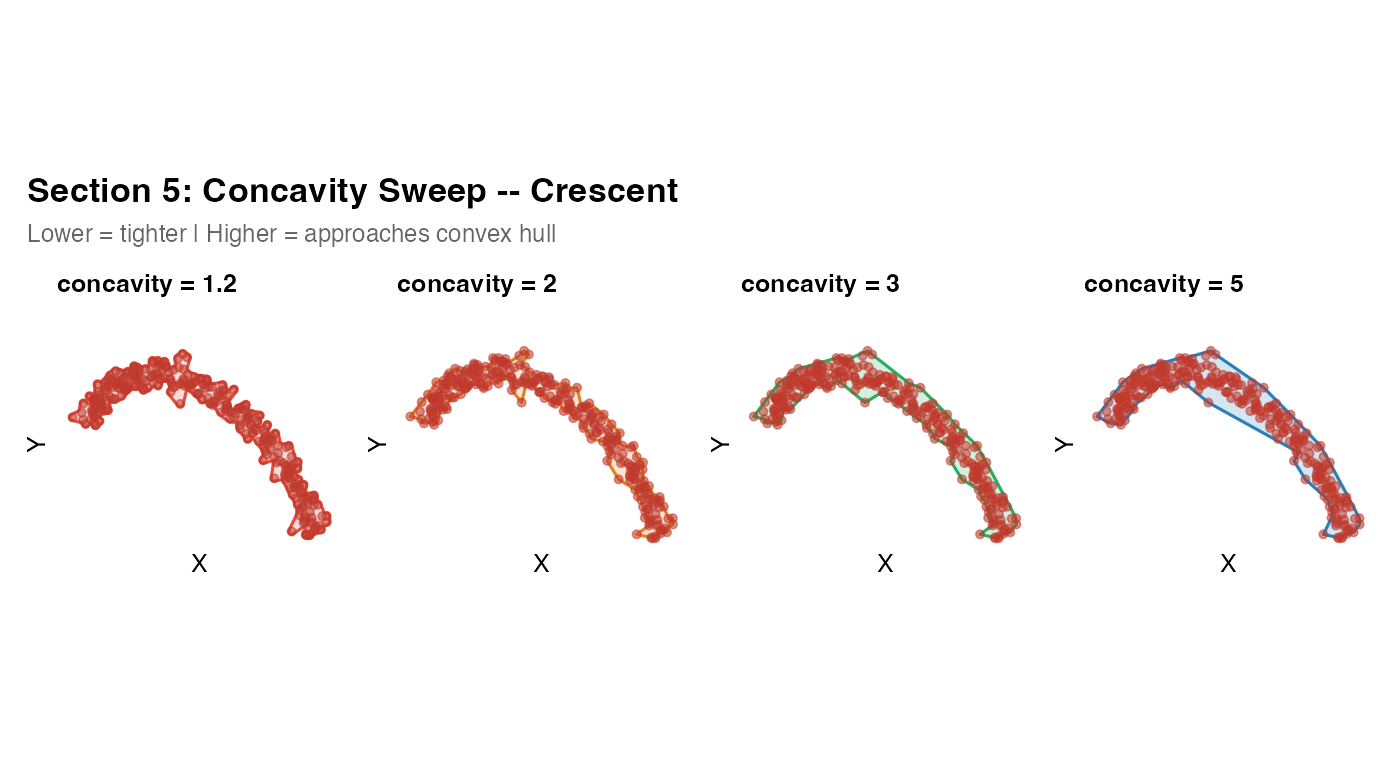
What to look for: At concavity = 1.2,
the hull cuts tightly around the arc. At concavity = 5, it
is nearly convex and fills the interior. The “correct” value for real
tissue is the one that matches a human drawing of the tissue boundary.
For a crescent-shaped airway section, 1.2--1.5 is usually
appropriate.
Section 6 – Buffer and smoothing
After fitting, two post-processing options are available:
-
buffer_dist: expands the mask outward by this distance in coordinate units usingsf::st_buffer(). Useful when you want a small safety margin to ensure all cells fall inside the mask, or when points are sparse near the boundary. -
smooth_mask: appliessf::st_simplify()to reduce the vertex count and round jagged boundary segments. Controlled bysmooth_tolerance(default: 1% of bounding box diagonal).
r <- sqrt(runif(250, 0.4^2, 1^2))
ang <- runif(250, 0, 2 * pi)
coords_ring <- data.frame(x = r * cos(ang) * 5,
y = r * sin(ang) * 5)
m6a <- fit_spatial_mask(coords_ring, method = "concave", concavity = 2,
buffer_dist = 0, smooth_mask = FALSE, verbose = FALSE)
m6b <- fit_spatial_mask(coords_ring, method = "concave", concavity = 2,
buffer_dist = 0.3, smooth_mask = FALSE, verbose = FALSE)
m6c <- fit_spatial_mask(coords_ring, method = "concave", concavity = 2,
buffer_dist = 0.3, smooth_mask = TRUE,
smooth_tolerance = 0.15, verbose = FALSE)
plot_mask(m6a, coords_ring,
title = "6a. Raw -- no buffer, no smooth") +
plot_mask(m6b, coords_ring,
title = "6b. Buffer = 0.3",
fill = "#27ae60", border = "#1e8449") +
plot_mask(m6c, coords_ring,
title = "6c. Buffer + smooth",
subtitle = "smooth_tolerance = 0.15",
fill = "#8e44ad", border = "#6c3483") +
plot_annotation(
title = "Section 6: Buffer and Smoothing",
theme = theme(plot.title = element_text(face = "bold", size = 13)))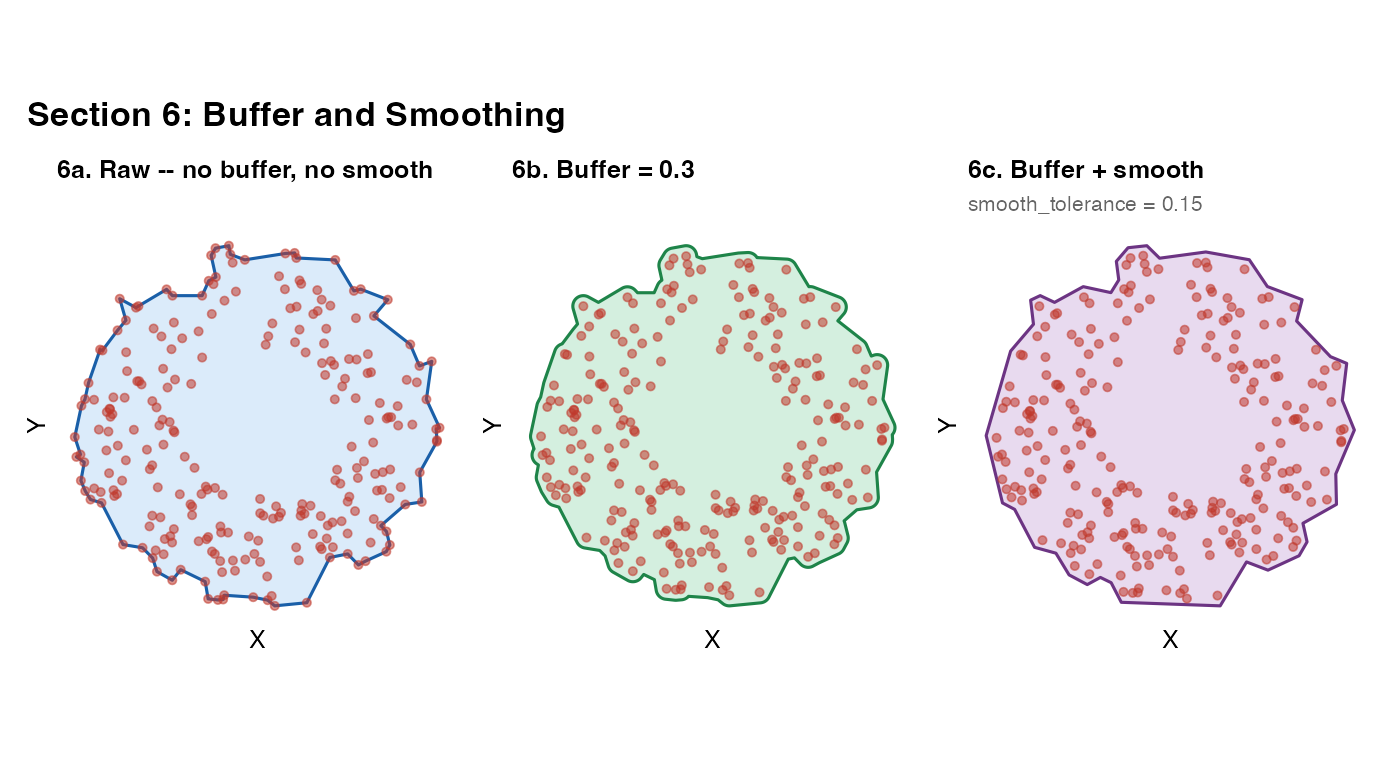
When to use each:
-
buffer_dist: use when downstream point-in-mask tests are failing for points you know should be inside (e.g. points right on the boundary). A value of 1–5% of the domain width is usually safe. -
smooth_mask: use when you want a visually clean boundary for publication figures, or when a jagged boundary is causing downstream geometric operations to be slow or numerically unstable. Be careful: largesmooth_tolerancevalues can erase real tissue features.
Section 7 – Topologically complex cases: holes and islands
The "raster" method is the recommended approach for
masks that need to represent holes (vessel lumens,
necrotic cores, tissue tears) or disconnected islands.
It is also the fastest method for large point clouds (100k+ cells).
How the raster method works
-
Bin all points onto a regular
raster_resolution x raster_resolutiongrid. -
Convolve the binary occupancy grid with a 2D
isotropic Gaussian of width
raster_sigma(in coordinate units). This is equivalent to placing a Gaussian “blob” at every point and summing – the value at each cell represents how much point density is nearby. -
Threshold at
raster_threshold x max(field). Cells above threshold are “inside”; cells below are “outside”. -
Build one axis-aligned rectangle per inside cell,
then dissolve all rectangles with
sf::st_union(). GEOS handles the topology automatically – holes and islands emerge from the geometry without any ring-winding logic. - Smooth the staircase boundary with a morphological close (buffer out then in by ~half the cell diagonal).
Key tuning parameters
| Parameter | Effect |
|---|---|
raster_sigma (larger) |
Holes fill in, islands merge, boundary smooths |
raster_sigma (smaller) |
Fine holes and gaps preserved |
raster_threshold (larger) |
Mask shrinks; requires denser local coverage |
raster_threshold (smaller) |
Mask grows; accepts sparse regions |
raster_resolution (larger) |
Finer grid; more detail; slower union step |
Units note:
raster_sigmais in coordinate units (same as x and y). If your tissue coordinates span 0–10, araster_sigmaof 0.25 means each Gaussian blob has a standard deviation of 0.25 tissue units.NULLsets sigma automatically to 3% of the domain width.
7a – Donut: single interior void
r_d <- sqrt(runif(800, 2^2, 5^2))
th_d <- runif(800, 0, 2 * pi)
coords_donut <- data.frame(x = r_d * cos(th_d),
y = r_d * sin(th_d))
m7a_r <- fit_spatial_mask(coords_donut, method = "raster",
raster_resolution = 256L, raster_sigma = 0.25, raster_threshold = 0.2,
verbose = FALSE)
m7a_c <- fit_spatial_mask(coords_donut, method = "convex", verbose = FALSE)
plot_mask(m7a_r, coords_donut,
title = "7a-i. Raster -- hole detected",
subtitle = "sigma = 0.25 coord units, threshold = 0.2") +
plot_mask(m7a_c, coords_donut,
title = "7a-ii. Convex -- hole filled",
subtitle = "Convex hull cannot represent holes",
fill = "#e74c3c", border = "#922b21") +
plot_annotation(
title = "Donut: Single Interior Void",
theme = theme(plot.title = element_text(face = "bold", size = 13)))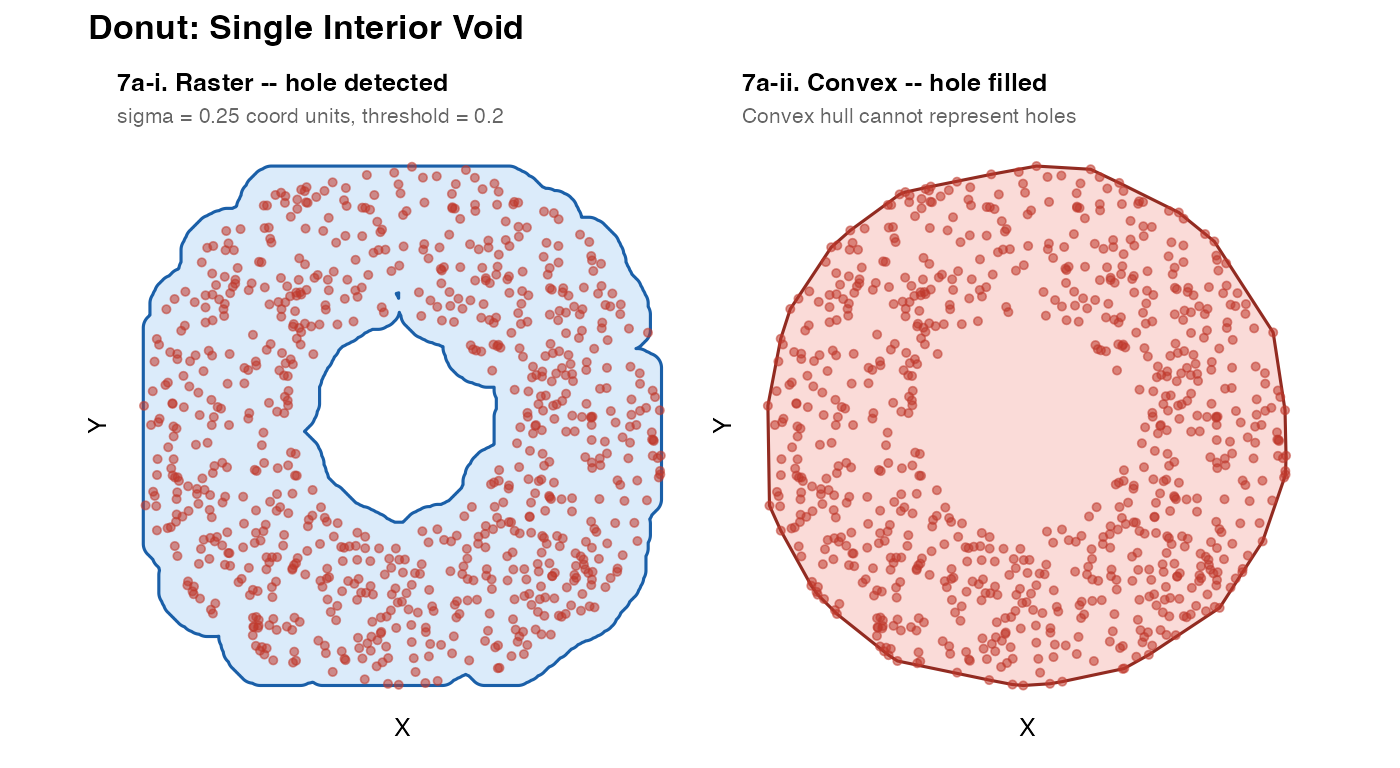
The raster method naturally detects the empty centre as a region of low convolved density, below the threshold. The convex hull cannot ever represent a hole – it is by definition the smallest convex polygon containing all the points.
7b – Swiss cheese: multiple holes
.circle_mat <- function(cx, cy, r, n = 100) {
th <- seq(0, 2 * pi, length.out = n)
mat <- cbind(cx + r * cos(th), cy + r * sin(th))
rbind(mat, mat[1L, ])
}
hdefs <- list(
list(cx = 2.5, cy = 7.5, r = 1.1),
list(cx = 7.0, cy = 6.5, r = 1.4),
list(cx = 4.0, cy = 2.5, r = 0.9),
list(cx = 8.0, cy = 2.0, r = 1.2)
)
sample_in_polygon <- function(poly_sf, n, max_tries = 20) {
bb <- sf::st_bbox(poly_sf)
mu <- sf::st_union(poly_sf)
pts <- data.frame(x = numeric(0), y = numeric(0))
for (i in seq_len(max_tries)) {
if (nrow(pts) >= n) break
cnd <- data.frame(x = runif(n * 8, bb["xmin"], bb["xmax"]),
y = runif(n * 8, bb["ymin"], bb["ymax"]))
ok <- as.logical(sf::st_within(
sf::st_as_sf(cnd, coords = c("x","y")), mu, sparse = FALSE)[, 1L])
pts <- rbind(pts, cnd[ok, ])
}
pts[seq_len(min(n, nrow(pts))), ]
}
outer_sq <- sf::st_sfc(sf::st_polygon(list(
rbind(c(0,0), c(10,0), c(10,10), c(0,10), c(0,0)))))
hcircles <- lapply(hdefs, function(h)
sf::st_polygon(list(.circle_mat(h$cx, h$cy, h$r))))
swiss_region <- sf::st_difference(outer_sq,
sf::st_sfc(sf::st_union(sf::st_sfc(hcircles))))
coords_swiss <- sample_in_polygon(swiss_region, n = 1500)
m7b_r <- fit_spatial_mask(coords_swiss, method = "raster",
raster_resolution = 256L, raster_sigma = 0.25, raster_threshold = 0.2,
verbose = FALSE)
m7b_c <- fit_spatial_mask(coords_swiss, method = "concave",
concavity = 1.8, verbose = FALSE)
plot_mask(m7b_r, coords_swiss,
title = "7b-i. Raster -- all 4 holes detected",
subtitle = "sigma = 0.25, threshold = 0.2") +
plot_mask(m7b_c, coords_swiss,
title = "7b-ii. Concave -- holes filled",
subtitle = "Concave hull also cannot represent holes",
fill = "#e74c3c", border = "#922b21") +
plot_annotation(
title = "Swiss Cheese: Multiple Interior Holes",
theme = theme(plot.title = element_text(face = "bold", size = 13)))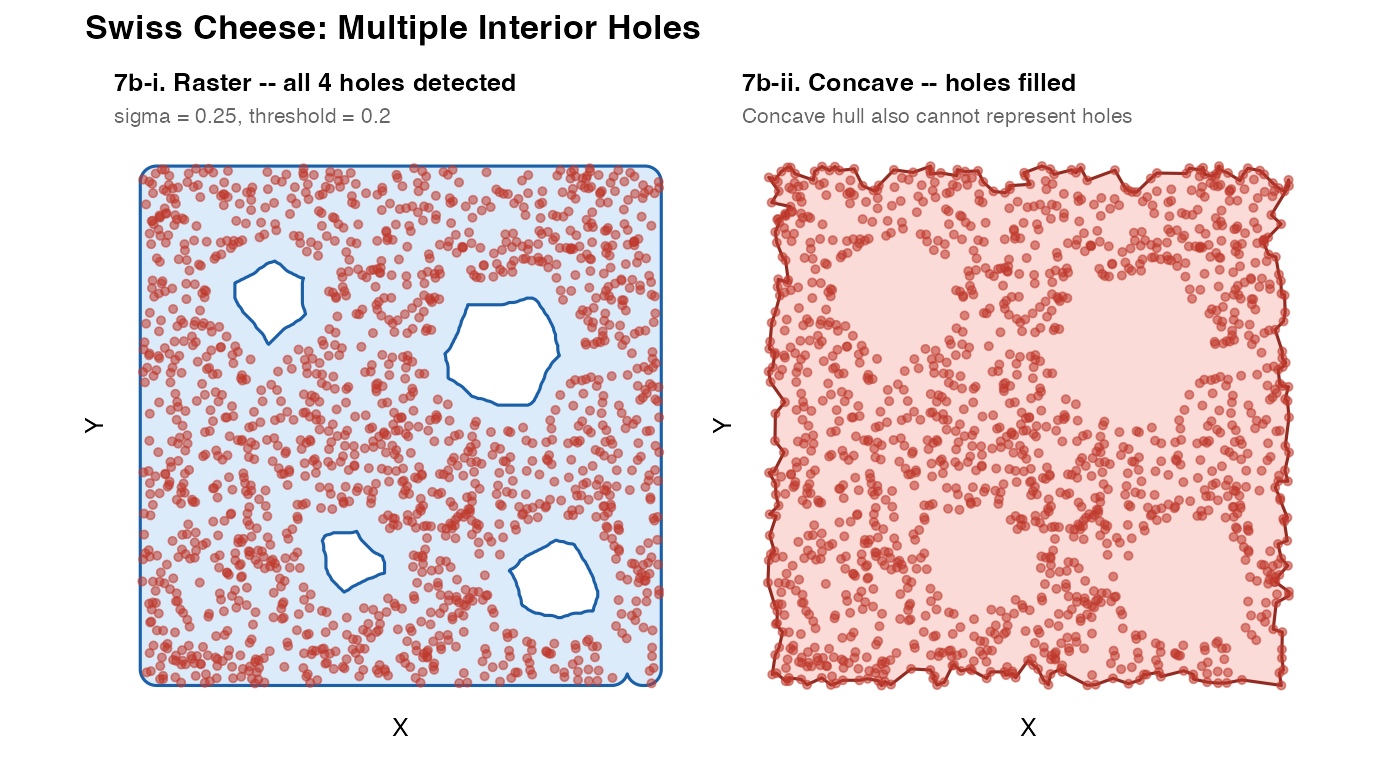
7c – Archipelago: disconnected islands
mk_isl <- function(n, cx, cy, rx, ry, ns = 0.15) {
th <- runif(n, 0, 2 * pi)
r <- sqrt(runif(n))
data.frame(x = cx + r * rx * cos(th) + rnorm(n, 0, ns),
y = cy + r * ry * sin(th) + rnorm(n, 0, ns))
}
coords_arch <- rbind(mk_isl(300, 0, 0, 3, 2),
mk_isl(300, 9, 4, 2, 3),
mk_isl(300, 5, -7, 2.5, 1.5))
m7c_r <- fit_spatial_mask(coords_arch, method = "raster",
raster_resolution = 256L, raster_sigma = 0.4, raster_threshold = 0.18,
verbose = FALSE)
m7c_c <- fit_spatial_mask(coords_arch, method = "convex", verbose = FALSE)
n_geom <- length(sf::st_cast(m7c_r, "POLYGON"))
plot_mask(m7c_r, coords_arch,
title = sprintf("7c-i. Raster -- %d separate polygons", n_geom),
subtitle = "Each island detected independently") +
plot_mask(m7c_c, coords_arch,
title = "7c-ii. Convex -- islands bridged",
subtitle = "Convex hull merges all islands into one",
fill = "#e74c3c", border = "#922b21") +
plot_annotation(
title = "Archipelago: Disconnected Islands",
theme = theme(plot.title = element_text(face = "bold", size = 13)))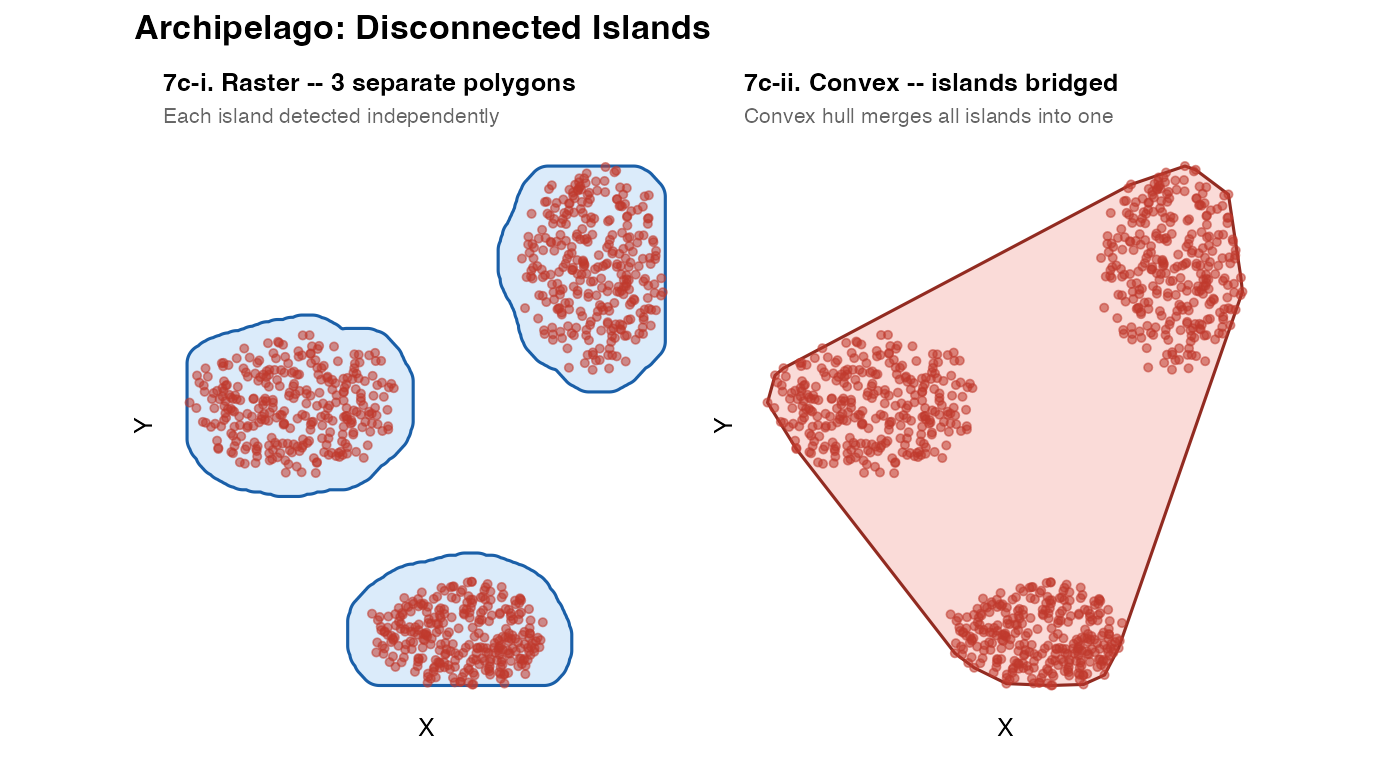
Disconnected islands emerge automatically from the GEOS union step:
if two groups of “on” cells are not touching, they become separate
polygons within the same sfc object. The number of distinct
polygons is accessible via
length(sf::st_cast(mask, "POLYGON")).
7d – Sigma sensitivity sweep
This is the most important tuning exercise. Run it on your own data
before finalising raster_sigma.
sig_vals <- c(0.1, 0.2, 0.4, 0.7)
sig_cols <- c("#e74c3c", "#e67e22", "#27ae60", "#2980b9")
sig_pls <- mapply(function(sv, col) {
m <- fit_spatial_mask(coords_swiss, method = "raster",
raster_resolution = 256L, raster_sigma = sv,
raster_threshold = 0.2, verbose = FALSE)
nh <- max(0L, length(sf::st_cast(m, "POLYGON")) - 1L)
plot_mask(m, coords_swiss,
title = paste0("sigma = ", sv, " coord units"),
subtitle = paste0(nh, " hole(s) detected"),
fill = col, border = col, pt_size = 0.5)
}, sig_vals, sig_cols, SIMPLIFY = FALSE)
wrap_plots(sig_pls, nrow = 1) +
plot_annotation(
title = "7d: raster_sigma Sensitivity -- Swiss Cheese",
subtitle = "Small sigma -> holes preserved | Large sigma -> holes filled",
theme = theme(plot.title = element_text(face = "bold", size = 13),
plot.subtitle = element_text(size = 9, color = "grey40")))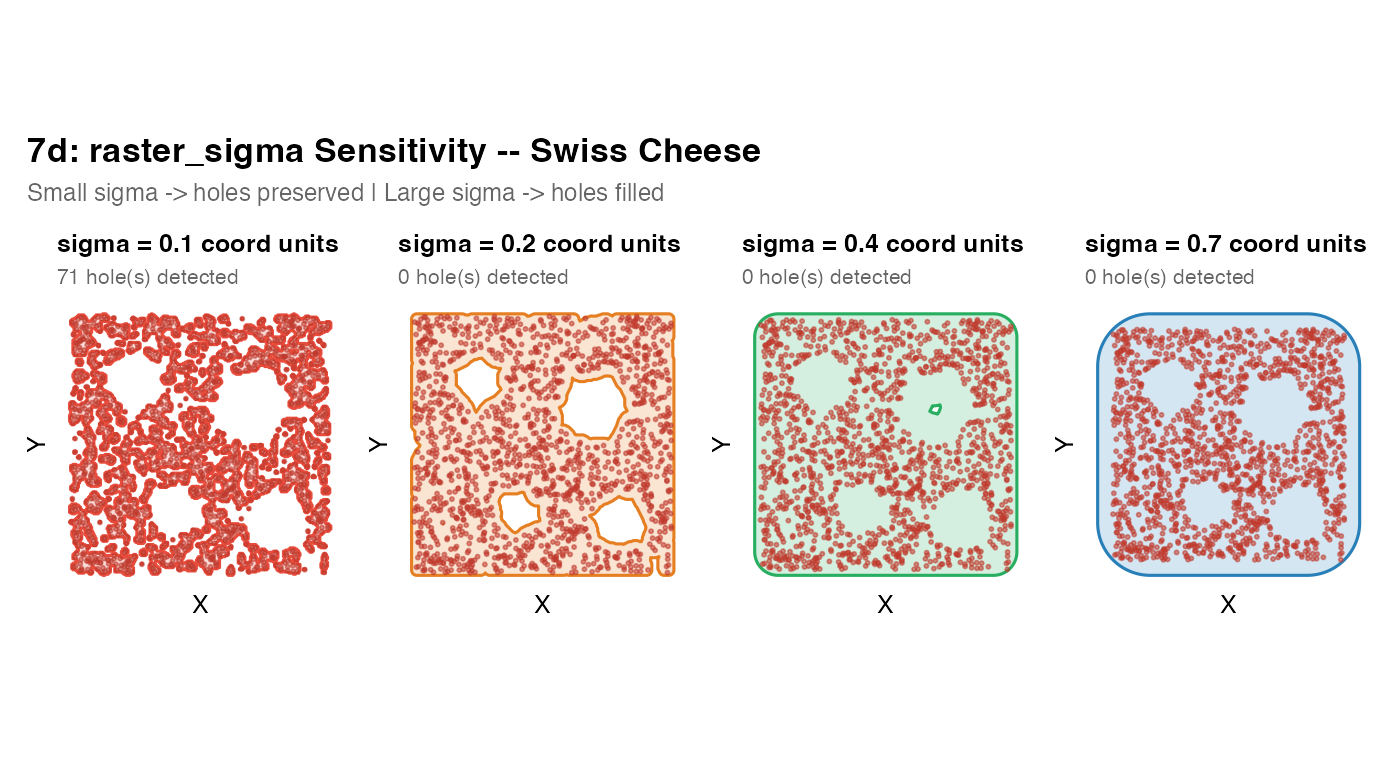
Tuning strategy:
- Start with
raster_sigma = NULL(auto = 3% of domain width) - Inspect the result; note whether real voids are detected
- Decrease sigma if holes are being filled that you expect to be present
- Increase sigma if the mask is too jagged or breaks into spurious fragments where there is actually continuous tissue
- Adjust
raster_thresholdto expand or contract the mask after sigma is set
7e – Nested crescent voids
mk_cres <- function(cx, cy, ro, ri, dx, dy, n = 100) {
moc <- .circle_mat(cx, cy, ro, n)
mic <- .circle_mat(cx+dx, cy+dy, ri, n)
oc <- sf::st_polygon(list(moc))
ic <- sf::st_polygon(list(mic))
sf::st_sfc(sf::st_difference(sf::st_sfc(oc), sf::st_sfc(ic)))
}
th_od <- seq(0, 2 * pi, length.out = 200)
mat_od <- cbind(10 * cos(th_od), 10 * sin(th_od))
mat_od <- rbind(mat_od, mat_od[1L, ])
outer_d <- sf::st_sfc(sf::st_polygon(list(mat_od)))
cres1 <- mk_cres(-2, 2, 2.5, 2, 0.9, -0.2)
cres2 <- mk_cres( 3, -3, 2.2, 1.8, -0.7, 0.4)
nest_region <- sf::st_difference(outer_d, sf::st_union(c(cres1, cres2)))
coords_nested <- sample_in_polygon(nest_region, n = 2500)
m7e_fine <- fit_spatial_mask(coords_nested, method = "raster",
raster_resolution = 256L, raster_sigma = 0.4, raster_threshold = 0.18,
verbose = FALSE)
m7e_coarse <- fit_spatial_mask(coords_nested, method = "raster",
raster_resolution = 256L, raster_sigma = 1.0, raster_threshold = 0.18,
verbose = FALSE)
plot_mask(m7e_fine, coords_nested,
title = "7e-i. Fine sigma = 0.4",
subtitle = "Crescent voids preserved") +
plot_mask(m7e_coarse, coords_nested,
title = "7e-ii. Coarse sigma = 1.0",
subtitle = "Narrow voids filled in",
fill = "#8e44ad", border = "#6c3483") +
plot_annotation(
title = "Nested Crescent Voids: sigma controls void resolution",
subtitle = "The minimum detectable void width is approximately 2 x raster_sigma",
theme = theme(plot.title = element_text(face = "bold", size = 13),
plot.subtitle = element_text(size = 9, color = "grey40")))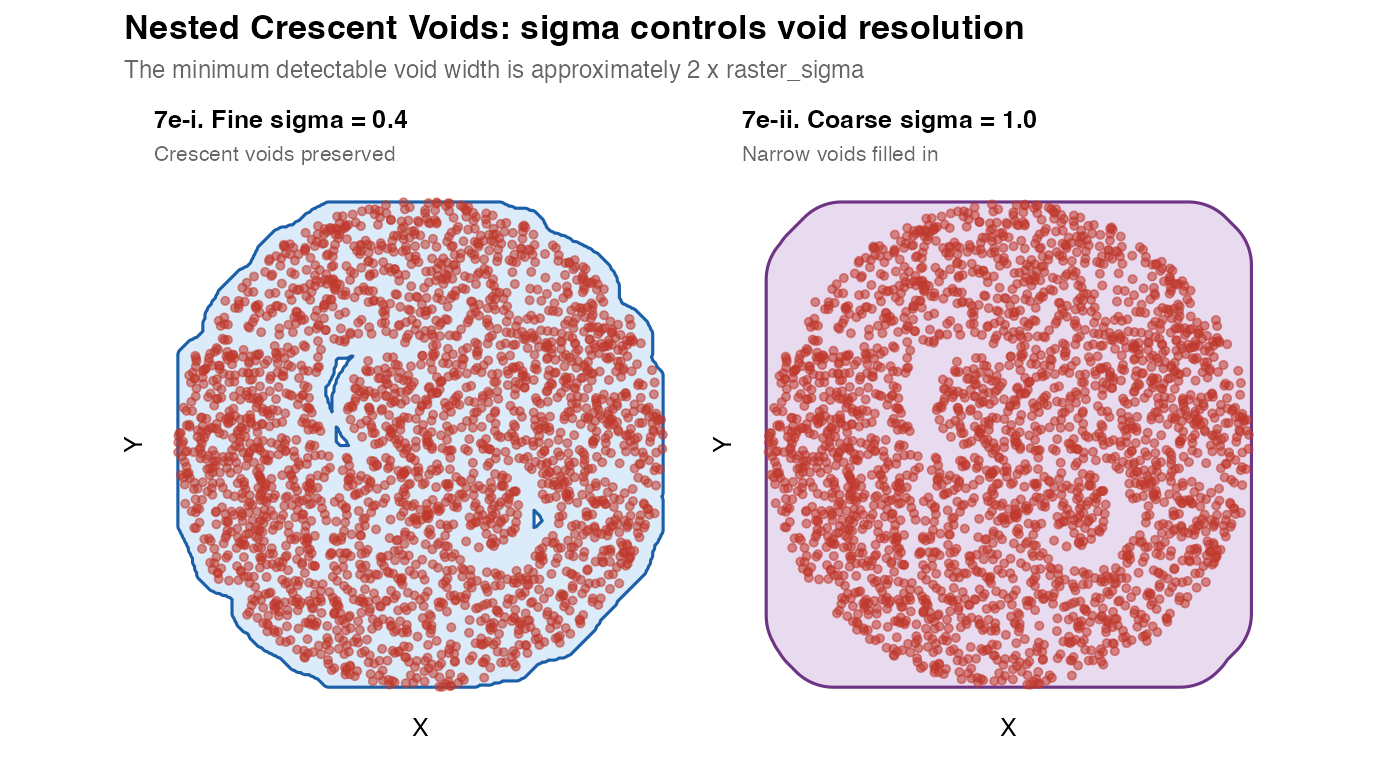
Key principle: The minimum resolvable void has a width of approximately
2 x raster_sigma. Voids narrower than this will be filled in by the Gaussian blur. If you need to resolve fine voids (e.g. individual capillary lumens), you must use a small sigma – but this may also make the mask jagged elsewhere.
7f – Scale test: 100,000 cells
The raster method is O(N^2) in the grid size but O(n) in the number
of input points (binning is a single tabulate() call). This
makes it practical for large datasets.
th100 <- seq(0, 2 * pi, length.out = 400)
r100 <- 12 + sin(5 * th100) + 0.4 * cos(11 * th100)
mat100 <- cbind(r100 * cos(th100), r100 * sin(th100))
mat100 <- rbind(mat100, mat100[1L, ])
outer100 <- sf::st_sfc(sf::st_polygon(list(mat100)))
make_ellipse_poly <- function(cx, cy, a, b, angle_deg, n = 80) {
th <- seq(0, 2 * pi, length.out = n)
ang <- angle_deg * pi / 180
xe <- a * cos(th); ye <- b * sin(th)
mat <- cbind(cx + xe * cos(ang) - ye * sin(ang),
cy + xe * sin(ang) + ye * cos(ang))
mat <- rbind(mat, mat[1L, ])
sf::st_polygon(list(mat))
}
holes100 <- sf::st_sfc(list(
make_ellipse_poly( 3, 5, 3, 1.5, 10),
make_ellipse_poly(-5, 2, 2.5, 2, -30),
make_ellipse_poly( 1, -6, 2, 3, 50)
))
reg100 <- sf::st_difference(outer100, sf::st_union(holes100))
coords_100k <- sample_in_polygon(reg100, n = 100000)
t_raster <- system.time(
m_100k <- fit_spatial_mask(coords_100k, method = "raster",
raster_resolution = 256L, raster_sigma = 0.6,
raster_threshold = 0.18, verbose = TRUE)
)## --------------------------------------------------
## TissueMask: spatial mask fitted
## Method : raster
## Points : 100000
## Sub-geometries : 1
## Bounding box : x [ -12.93 , 12.629 ] y [ -11.827 , 13.227 ]
## Area : 523.5807
## Buffer applied : 0
## --------------------------------------------------
sub <- coords_100k[sample(nrow(coords_100k), 5000), ]
plot_mask(m_100k, sub,
title = sprintf("7f. 100k cells -- raster (%.1f s)", t_raster["elapsed"]),
subtitle = "3 elliptic holes detected | 5,000 of 100,000 points shown",
pt_size = 0.3, pt_alpha = 0.2)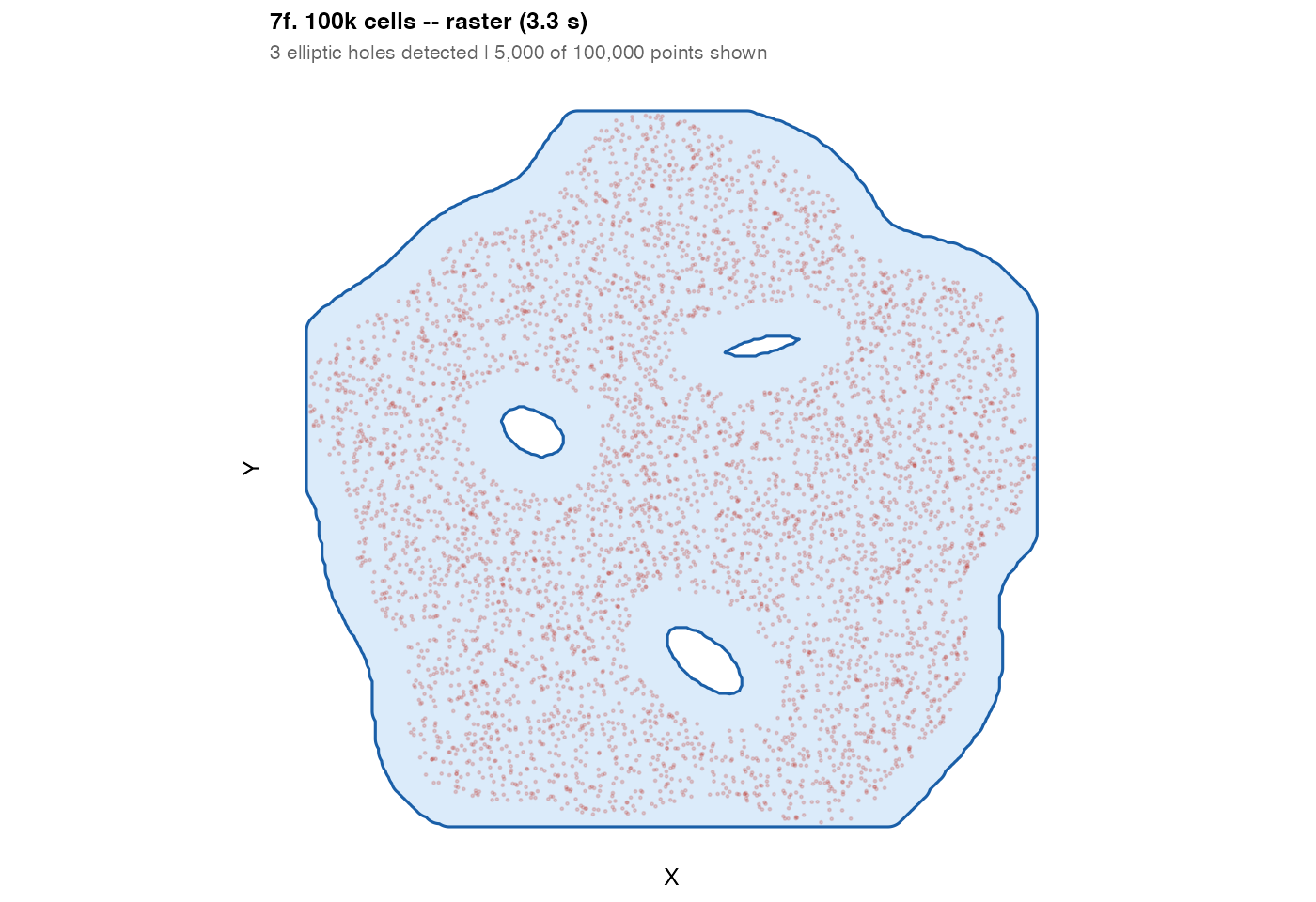
Section 8 – Validation: point containment
A properly fitted mask must contain all its input points. This table
checks every mask produced in this vignette. Any FAIL indicates that the
mask needs a larger buffer_dist or a lower
raster_threshold.
cases <- list(
list(label = "Circle/convex", mask = m1a, coords = coords_circle),
list(label = "Circle/concave", mask = m1b, coords = coords_circle),
list(label = "Circle/kde", mask = m1c, coords = coords_circle),
list(label = "Elongated/convex", mask = m2a, coords = coords_elong),
list(label = "Elongated/concave", mask = m2b, coords = coords_elong),
list(label = "Elongated/kde", mask = m2c, coords = coords_elong),
list(label = "L-shape/convex", mask = m3a, coords = coords_L),
list(label = "L-shape/conc1.5", mask = m3b, coords = coords_L),
list(label = "L-shape/conc3", mask = m3c, coords = coords_L),
list(label = "Multi/convex", mask = m4a, coords = coords_multi),
list(label = "Multi/concave", mask = m4b, coords = coords_multi),
list(label = "Multi/kde-lo", mask = m4c, coords = coords_multi),
list(label = "Multi/kde-hi", mask = m4d, coords = coords_multi),
list(label = "Ring/raw", mask = m6a, coords = coords_ring),
list(label = "Ring/buffer", mask = m6b, coords = coords_ring),
list(label = "Ring/smooth", mask = m6c, coords = coords_ring),
list(label = "Donut/raster", mask = m7a_r, coords = coords_donut),
list(label = "Swiss/raster", mask = m7b_r, coords = coords_swiss),
list(label = "Archipelago/raster", mask = m7c_r, coords = coords_arch),
list(label = "Nested/fine", mask = m7e_fine, coords = coords_nested),
list(label = "Nested/coarse", mask = m7e_coarse,coords = coords_nested),
list(label = "100k/raster", mask = m_100k, coords = coords_100k)
)
results <- do.call(rbind, lapply(cases, function(vc) {
pts <- sf::st_geometry(sf::st_as_sf(vc$coords, coords = c("x","y")))
ok <- sf::st_covered_by(pts, sf::st_union(vc$mask), sparse = FALSE)[, 1L]
n <- nrow(vc$coords)
data.frame(
Case = vc$label,
N = n,
Enclosed = sum(ok),
Pct = round(100 * sum(ok) / n, 2),
Result = ifelse(sum(ok) == n, "PASS", "FAIL"),
stringsAsFactors = FALSE
)
}))
knitr::kable(results, align = c("l","r","r","r","c"))| Case | N | Enclosed | Pct | Result |
|---|---|---|---|---|
| Circle/convex | 400 | 400 | 100 | PASS |
| Circle/concave | 400 | 400 | 100 | PASS |
| Circle/kde | 400 | 400 | 100 | PASS |
| Elongated/convex | 400 | 400 | 100 | PASS |
| Elongated/concave | 400 | 400 | 100 | PASS |
| Elongated/kde | 400 | 400 | 100 | PASS |
| L-shape/convex | 400 | 400 | 100 | PASS |
| L-shape/conc1.5 | 400 | 400 | 100 | PASS |
| L-shape/conc3 | 400 | 400 | 100 | PASS |
| Multi/convex | 450 | 450 | 100 | PASS |
| Multi/concave | 450 | 450 | 100 | PASS |
| Multi/kde-lo | 450 | 450 | 100 | PASS |
| Multi/kde-hi | 450 | 450 | 100 | PASS |
| Ring/raw | 250 | 250 | 100 | PASS |
| Ring/buffer | 250 | 250 | 100 | PASS |
| Ring/smooth | 250 | 250 | 100 | PASS |
| Donut/raster | 800 | 800 | 100 | PASS |
| Swiss/raster | 1500 | 1500 | 100 | PASS |
| Archipelago/raster | 900 | 900 | 100 | PASS |
| Nested/fine | 2500 | 2500 | 100 | PASS |
| Nested/coarse | 2500 | 2500 | 100 | PASS |
| 100k/raster | 100000 | 100000 | 100 | PASS |
Quick-start decision guide
Your data
|
+-- Simple convex blob? -----------------------> method = "convex"
|
+-- Non-convex (C-shape, arc, L-shape)?
| +-- No holes needed? -------------------> method = "concave"
| concavity: start at 2, lower to tighten
|
+-- Holes or islands expected?
| +-- n < 50,000? -----------------------> method = "raster"
| +-- n > 50,000? -----------------------> method = "raster" (still fine)
|
+-- Dense, smooth, no need for holes?
+-- KDE is an alternative -------------> method = "kde"
kde_threshold: start at 0.05
For method = "raster":
raster_sigma = NULL (auto) for a first pass
raster_sigma smaller: preserve fine voids
raster_sigma larger: fill gaps and smooth edges
raster_threshold = 0.15 is a reliable defaultSession information
## R version 4.5.2 (2025-10-31)
## Platform: aarch64-apple-darwin20
## Running under: macOS Sonoma 14.6.1
##
## Matrix products: default
## BLAS: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libBLAS.dylib
## LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
##
## locale:
## [1] en_US/en_US/en_US/C/en_US/en_US
##
## time zone: America/New_York
## tzcode source: internal
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] patchwork_1.3.2 ggplot2_4.0.1 sf_1.0-21 TissueMask_0.1.0
##
## loaded via a namespace (and not attached):
## [1] sass_0.4.10 generics_0.1.4 class_7.3-23 KernSmooth_2.23-26
## [5] digest_0.6.39 magrittr_2.0.4 evaluate_1.0.5 grid_4.5.2
## [9] RColorBrewer_1.1-3 fastmap_1.2.0 jsonlite_2.0.0 e1071_1.7-16
## [13] DBI_1.2.3 scales_1.4.0 isoband_0.3.0 textshaping_1.0.1
## [17] jquerylib_0.1.4 cli_3.6.5 rlang_1.1.7 units_0.8-7
## [21] withr_3.0.2 cachem_1.1.0 yaml_2.3.12 otel_0.2.0
## [25] concaveman_1.2.0 tools_4.5.2 parallel_4.5.2 dplyr_1.1.4
## [29] curl_7.0.0 vctrs_0.7.1 R6_2.6.1 proxy_0.4-27
## [33] lifecycle_1.0.5 classInt_0.4-11 V8_8.0.1 fs_1.6.6
## [37] htmlwidgets_1.6.4 MASS_7.3-65 ragg_1.4.0 pkgconfig_2.0.3
## [41] desc_1.4.3 pkgdown_2.1.3 bslib_0.10.0 pillar_1.11.1
## [45] gtable_0.3.6 glue_1.8.0 Rcpp_1.1.1 systemfonts_1.2.3
## [49] xfun_0.56 tibble_3.3.1 tidyselect_1.2.1 knitr_1.51
## [53] dichromat_2.0-0.1 farver_2.1.2 htmltools_0.5.9 rmarkdown_2.30
## [57] compiler_4.5.2 S7_0.2.1